A Designer Enzyme for Alternative Energy
June 20, 2013
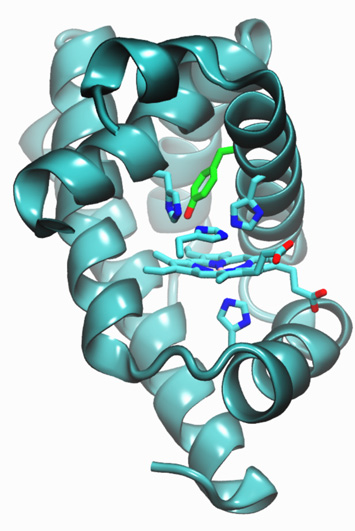
Ribbon diagram showing introduction of histadine and tyrosine residues into myoglobin, resulting in enzymes that reduce oxygen (O2)to water (H2O) with more than 1,000 turnovers and minimal release of reactive oxygen species
Imagine pulling energy out of thin air. Yi Lu and his colleagues are on that path, in a quest to find alternatives to fossil fuels. The team has designed an enzyme that can harvest the energy of atmospheric oxygen with high efficiency and long life. Their work is a big step in custom designing artificial enzymes for potential applications in alternative energy.
"Oxygen in air is abundant and cheap. Yet converting it to useful energy requires catalysts with high efficiency and stability," said Lu, a professor at the University of Illinois. He explained that nature's catalysts are among the most efficient, certainly better than most fuel-cell catalysts of today. A typical "native" catalyst is the enzyme heme copper oxidase, or HCO. But HCOs are too big, too expensive and too unstable for practical applications.
"We have designed an enzyme that is much smaller, cheaper and more stable than the native enzymes," said Lu.
In brief, they introduced one tyrosine and two histidine residues into myoglobin, which yielded an enzyme that catalyzed the reduction of oxygen to water with minimal release of reactive oxygen species and more than 1,000 turnovers – the number of times an enzyme can catalyze the same reaction over and over. Reactive oxygen species are incomplete intermediates for the oxygen reduction; they not only decrease the efficiency of the energy-conversion reaction, but also damage fuel-cell components.
According to Lu, designing enzymes, especially ones such as theirs that contain metal ions, is a real challenge. Most designed enzymes have relatively low turnovers and efficiency, making them difficult to use for applications. The researchers demonstrated the feasibility of designing enzymes with both high turnovers and efficiency.
From initial design of the enzymes to final confirmation of the design, obtaining high-resolution crystal structures is essential at every stage of this project, said Lu. That work has been done with Photon Sciences (PS) biologist Howard Robinson at Brookhaven National Laboratory's National Synchrotron Light Source (NSLS). Looking ahead, the research will continue at NSLS-II, expected to start commissioning at Brookhaven in 2014. Said Lu, "The new facility will offer even better capability to obtain crystal structures. It will also allow simultaneous study of our designed enzymes by both crystallography and single-crystal spectroscopy." The team has already started working toward that direction with Allen Orville, another PS biologist.
This current work on enzyme design was published in the journal Angewandte Chemie International Edition. The complete author list is: Kyle Miner, Arnab Mukherjee, Yi-Gui Gao, Eric Null, Igor Petrik, Xuan Zhao, Natasha Yeung, Howard Robinson and Yi Lu.
2013-4073 | INT/EXT | Newsroom