Computation and Data-Driven Discovery (C3D) Projects
Complex Modeling of Nanostructures
This project seeks to develop multimodal data analysis methods and software tools for the characterization of nanomaterials structures, by combining the analysis of multiple experimental inputs with numerical simulations. Accurate knowledge of the atomic structure is the foundation for theoretical understanding of the material’s behavior and is thus critical for design and guided development of new materials. Conventional crystallography can almost routinely determine the structure of bulk crystalline materials, as their long-range periodic order and high-symmetries of atom arrangements allow structure description with a few variables. The situation is however much more difficult for nanomaterials. Nanostructures have too few atoms to form a clear periodic order in their positions. A significant share of their atoms is at the surface where atom positions relax and distort. Nanoparticles thus adopt more complex, lower-symmetry arrangements than their bulk counterparts. They also produce noisier, less resolved X-ray diffraction patterns that are inadequate for solving their more complicated structures with more unknown variables. To overcome these difficulties, we combine multiple experimental inputs such as pair distribution functions (PDF), small angle scattering (SAS), local chemical constraints (rigid groups, allowed bond angle ranges) and theoretical energy calculations in a single optimization setup. The increased number of inputs that the solved structure has to compensate for the structure’s additional degrees of freedom and for the lower information content in the experimental data and thus facilitates more complete depiction of the structure, Fig. 1.
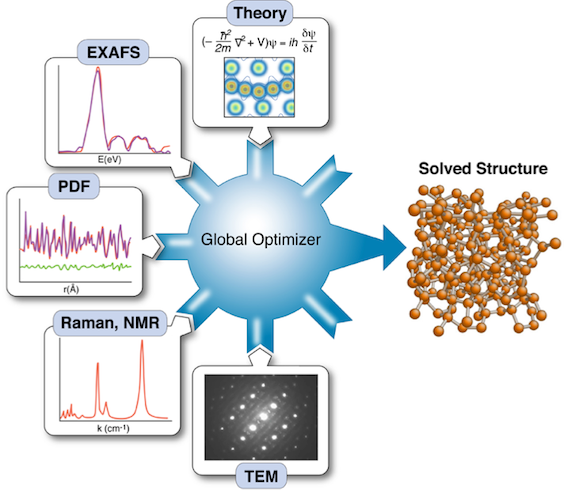
Fig. 1. Complex Modeling combines multiple experimental and theoretical inputs in a global optimizer to determine structure of a new material
To conduct such types of analysis we have developed an open-source software framework DiffPy-CMI (Complex Modeling Infrastructure). DiffPy-CMI is a collection of Python and C++ libraries that provide objects for diverse structure representations and operations on structure data, functions for simulating PDF, SAS, bond valence sums and overlap of empirical atom radii. Complex Modeling is achieved with a top-level utility for managing multiple data refinements and combining them in a single optimization problem. The object-oriented architecture of DiffPy-CMI aims to maximize code reuse and provide high flexibility so that simulations can be tweaked and tailored to the specifics of the studied material. The DiffPy-CMI codes are developed as open source software to promote sharing, verification and contributions from the community.
Publications
P. Juhás, C. L. Farrow, X. Yang, K. R. Knox and S. J. L. Billinge, Complex modeling: a strategy and software program for combining multiple information sources to solve ill posed structure and nanostructure inverse problems, Acta Crystallogr. A 71, 562-568 (2015).
Dragica Prill, Pavol Juhas, S. J. L. Billinge and Martin U. Schmidt, Solution and refinement of organic crystal structures by fitting to the atomic pair distribution function (PDF), Acta Crystallogr. A 72, 62-72 (2016).
Alexander N. Beecher, Xiaohao Yang, Joshua H. Palmer, Alexandra L. LaGrassa, Pavol Juhás, Simon J. L. Billinge and Jonathan S. Owen, Atomic structures and gram scale synthesis of three tetrahedral quantum dots, J. Am. Chem. Soc. 136, 10645-10653 (2014).
Christoper L. Farrow, Chenyang Shi, Pavol Juhás, Xiaogang Peng, and Simon J. L. Billinge, Robust structure and morphology parameters for CdS nanoparticles by combining small angle X-ray scattering and atomic pair distribution function data in a complex modeling framework, J. Appl. Crystallogr. 47, 561-565 (2014).
Kirsten M. Ø. Jensen, Mogens Christensen, Pavol Juhás, Christoffer Tyrsted, Espen D. Bøjesen, Nina Lock, Simon J. L. Billinge, and Bo B. Iversen, Revealing the mechanisms behind SnO2 nanoparticle formation and growth during hydrothermal synthesis: an in situ total scattering study, J. Am. Chem. Soc. 134, 6785 (2012).
Online References
- http://www.diffpy.org/
- http://github.com/diffpy/
- Google Group diffpy-users




