Modeling Plant Metabolism to Optimize Oil Production
Computational studies aim to increase use of plant oils as renewable resource
July 26, 2011
 enlarge
enlarge
Developing embryos after being excised from a growing rapeseed plant. The embryos accumulate seed oils which represent the most energy-dense form of biologically stored sunlight, and have great potential as renewable resources for fuel and industrial chemicals.
UPTON, NY — Scientists at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory have developed a computational model for analyzing the metabolic processes in rapeseed plants — particularly those related to the production of oils in their seeds. Their goal is to find ways to optimize the production of plant oils that have widespread potential as renewable resources for fuel and industrial chemicals.
The model, described in two “featured articles” in the August 1, 2011, issue of the Plant Journal (now available online*), may help to identify ways to maximize the conversion of carbon to biomass to improve the production of plant-derived biofuels.
“To make efficient use of all that plants have to offer in terms of alternative energy, replacing petrochemicals in industrial processes, and even nutrition, it’s essential that we understand their metabolic processes and the factors that influence their composition,” said Brookhaven biologist Jorg Schwender, who led the development of the model with postdoctoral research associate Jordan Hay.
In the case of plant oils, the scientists’ attention is focused on seeds, where oils are formed and accumulated during development. “This oil represents the most energy-dense form of biologically stored sunlight, and its production is controlled, in part, by the metabolic processes within developing seeds,” Schwender said.
One way to study these metabolic pathways is to track the uptake and allotment of a form of carbon known as carbon-13 as it is incorporated into plant oil precursors and the oils themselves. But this method has limits in the analysis of large-scale metabolic networks such as those involved in apportioning nutrients under variable physiological conditions.
“It’s like trying to assess traffic flow on roads in the United States by measuring traffic flow only on the major highways,” Schwender said.
To address these more complex situations, the Brookhaven team constructed a computational model of a large-scale metabolic network of developing rapeseed (Brassica napus) embryos, based on information mined from biochemical literature, databases, and prior experimental results that set limits on certain variables. The model includes 572 biochemical reactions that play a role in the seed’s central metabolism and/or seed oil production, and incorporates information on how those reactions are grouped together and interact.
 enlarge
enlarge
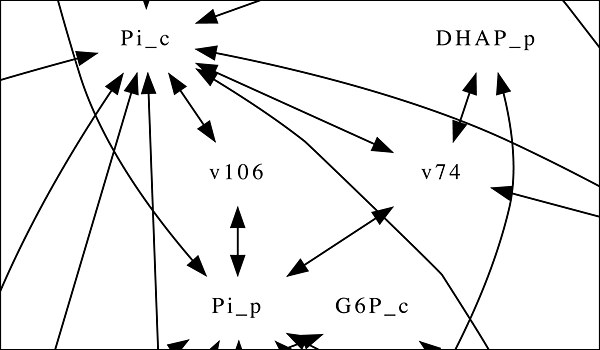
A network illustrating some of the reactions and chemical pathways involved in oil production in rapeseed plants. By modeling these interacting pathways, scientists may find ways to optimize plant oil production so the oils can be used as fuels or raw materials for industrial processes.
The scientists first tested the validity of the model by comparing it to experimental results from carbon-tracing studies for a relatively simple reaction network — the big-picture view of the metabolic pathways analogous to the traffic on U.S. highways. At that big-picture level, results from the two methods were largely consistent, providing validation for both the computer model and the experimental technique, while identifying a few exceptions that merit further exploration.
The scientists then used the model to simulate more complicated metabolic processes under varying conditions — for example, changes in oil production or the formation of oil precursors in response to changes in available nutrients (such as different sources of carbon and nitrogen), light conditions, and other variables.
“This large-scale model is a much more realistic network, like a map that represents almost every street,” Schwender said, “with computational simulations to predict what’s going on.” Continuing the traffic analogy, he said, “We can now try to simulate the effect of ‘road blocks’ or where to add new roads to most effectively eliminate traffic congestion.”
The model also allows the researchers to assess the potential effects of genetic modifications (for example, inactivating particular genes that play a role in plant metabolism) in a simulated environment. These simulated “knock-out” experiments gave detailed insights into the potential function of alternative metabolic pathways — for example, those leading to the formation of precursors to plant oils, and those related to how plants respond to different sources of nitrogen.
“The model has helped us construct a fairly comprehensive overview of the many possible alternative routes involved in oil formation in rapeseed, and categorize particular reactions and pathways according to the efficiency by which the organism converts sugars into oils. So at this stage, we can enumerate, better than before, which genes and reactions are necessary for oil formation, and which make oil production most effective,” Schwender said.
The researchers emphasize that experimentation will still be essential to further elucidating the factors that can improve plant oil production. “Any kind of model is a largely simplified representation of processes that occur in a living plant,” Schwender said. “But it provides a way to rapidly assess the relative importance of multiple variables and further refine experimental studies. In fact, we see our model and experimental methods such as carbon tracing as complementary ways to improve our understanding of plants’ metabolic pathways.”
The scientists are already incorporating information from this study that will further refine the model to increase its predictive power, as well as ways to extend and adapt it for use in studying other plant systems.
This work was supported by the DOE Office of Science.
2011-11308 | INT/EXT | Newsroom