CFN User Spotlight: Laura Fabris Develops Nanoparticle-Based Tags to Detect Cancer and Viruses at the Single-Cell Level
interview with a CFN user
April 24, 2018

Laura Fabris in a lab in the School of Engineering at Rutgers. Credit: Kate Woodside, Rutgers
How can we detect cancer and viruses with high sensitivity? Physical chemist Laura Fabris—an associate professor in the Materials Science and Engineering Department at Rutgers, the State University of New Jersey, and principal investigator of the Fabris NanoBio Group—is addressing this very question. Her research focuses on the synthesis, functionalization, characterization, and application of tiny metallic nanoparticles. These “plasmonic” nanoparticles greatly enhance the intensity of the signals produced by a surface-sensitive technique for detecting molecules. Fabris and her research group use the transmission electron microscopes at the Center for Functional Nanomaterials (CFN)—a U.S. Department of Energy Office of Science User Facility at Brookhaven National Laboratory—to visualize the nanoparticles and understand how to improve their morphology to improve clinical diagnoses.
Plasmonics is the study of how free electrons at the surface of a material are excited by light to have collective charge oscillations. What are plasmonic nanoparticles, and why are they of interest in biomedicine?
Traditionally, plasmonic nanoparticles are small metal particles that absorb light at different wavelengths depending on their properties—for example, gold absorbs light at 520 nanometers and silver at 400 nanometers—and amplify this light like a lens or concentrator does. Their strong interaction with light occurs because the electrons on the surface oscillate coherently with the light hitting them, in response generating an electric field that is locally much more intense than the original. This intense field can be taken advantage of downstream to increase the detection limit in a chemical analysis technique called Raman spectroscopy, which is used to identify the presence of biomolecules.
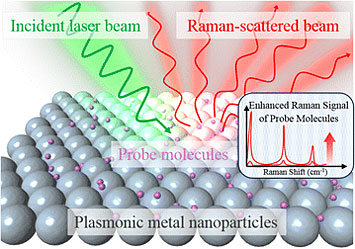
Plasmonic nanoparticles can be used to magnify the Raman scattering signal from molecules of interest by several orders of magnitude. Source: Nano Convergence, 2016, 3: 18.
My group focuses on understanding how the light-matter interactions change with the composition, shape, and size of the nanoparticles. For example, when electromagnetic fields interact with these metal nanoparticles, there is a high localization of the electric field in places with lots of curvature (corners, edges, tips). So if you can generate particles with lots of spiky regions, then there will be many areas—“hot spots”—where the electric field is enhanced. The electric field can be amplified anywhere from 1000 to 10,000 times, and this amplification is reflected downstream when we measure the intensity of the Raman signals. When molecules of interest are placed near the surface of these hot spots, the Raman signals can be enhanced by 10 or 11 orders of magnitude. This enhancement is sufficient for detecting single molecules.
My group has synthesized gold nanostars—spherical cores with many spiky tips like a sea urchin—for this very reason. A lot of discovery science in the growth of nanoparticles is often done with gold because it is much more stable as compared to some other metals. Silver and aluminum oxidize very readily, and copper does not absorb light as intensely and its shape is harder to tune. In terms of applications, gold is of interest because it is not cytotoxic—we can use it in cells without killing them.
How does Raman spectroscopy work? What information does it provide, and why does the detection limit need to be increased?
Raman spectroscopy is a technique used to measure molecular vibrations, providing a fingerprint of chemical bonds that are present in molecules. In this technique, light from a laser beam is directed at a sample. After molecules absorb this radiation, they are excited to a very short-lived, highly excited state. When the molecules relax to a lower energy level, they emit (scatter) light, which is collected by a detector that records the intensity. Any change in energy between the incident and scattered light is associated with specific molecular vibrations.
Because each molecule has a unique vibration frequency, Raman scattering can be used to determine a material’s chemical composition and crystal structure. This technique, which has been around for more than 100 years, works great for high concentrations of materials but it does not have the sensitivity to detect materials at low concentrations. For example, the concentration of proteins and other biomarkers in our body are too low for detection by regular Raman scattering.
But in 1977, scientists understood that if a molecule of interest happens to be in close proximity to plasmonic nanoparticles, then the intensity of the Raman signal gets amplified. This phenomenon is due to the enhanced scattered electric field that results when the nanoparticles interact with the electromagnetic radiation from the laser. Molecules adsorbed onto the surfaces of plasmonic structures experience this strongly enhanced field, and the end effect is that the Raman scattering signal obtained from these adsorbed molecules is a lot more intense. This technique is called surface-enhanced Raman scattering, or SERS.
How are you applying SERS in your research?
My group is looking for ways to monitor biologically relevant events that could not be monitored in other ways. One area we are interested in applying SERS to is oncology, to identify and image individual cancer cells and understand how they change as the disease progresses or in response to treatment. MRI [magnetic resonance imaging], a commonly used tool in cancer diagnosis, has a resolution limit of one millimeter. Thousands of cells exist in a cubic millimeter. We need to be able to see individual cancer cells to catch the disease before its growth becomes uncontrollable. SERS is ideally suited for this task because it is very concentration-sensitive and space-resolution sensitive.
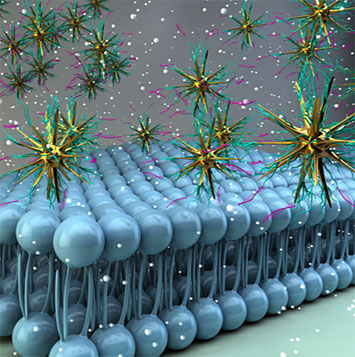

When gold nanostars are properly functionalized, they can selectively recognize cancerous cells and penetrate their membrane.
My group has developed SERS tags—a nanoparticle or nanoparticle assembly that is made of gold (so we can put it in cells), has a specific shape to ensure the field (and thus Raman signal) is enhanced a lot, and carries molecules that uniquely recognize cancer cells. When a cell is diseased, it has a high number of proteins in its membrane that are typical of cancer. On the Raman-active nanoparticles, we put antibodies or small peptides that recognize these proteins. Then, when the nanoparticles are added to a cell culture or the bloodstream, they specifically stick to these proteins. If we know the nanoparticle goes to a specific cell that we want to image and see that the enhanced Raman signal is coming from the same place where the cell is, we indirectly know that cell is cancerous.
This approach has been picking up in the last few years by various groups, including those at Memorial Sloan Kettering Cancer Center in New York City and Rutgers Cancer Institute of New Jersey. I am currently working with Dr. Isaac Kim, an oncologist at Rutgers Cancer Institute, who provides tissues from his prostate cancer patients. Having access to these tissues will help us better design nanoparticles to specifically target those cancer cells and correlate the information that the nanoparticles give us to disease progression.
The Fabris NanoBio Group consists of chemists, materials scientists, physicists, biomedical engineers, and electrical engineers who are developing new imaging tools to understand how cells evolve as a result of disease progression or treatment. This video explains how the group uses gold nanoparticles and surface-enhanced Raman scattering to assess the health of a single cell. Credit: Rutgers University Watch Video
Another area we are focusing on is virology—how can we detect and monitor viral mutations at the single-cell level with the SERS tags? I am collaborating with researchers from NC State University, the University of Illinois at Urbana-Champaign, Duke University, and Montana State University through a grant from the Defense Advanced Research Projects Agency (DARPA) to understand how the influenza virus penetrates a cell and under what conditions it is induced to mutate.
The flu and viruses in general have many different strains and mutate extremely rapidly, making them moving targets. How do you plan to detect and monitor something that is always changing?
The influenza virus has eight different segments. Each segment has one highly mutated region and two highly conserved regions that never mutate. With our particles, we are currently targeting the conserved regions to make sure that our tool can effectively identify the viral particles.
Right now, it is impossible to recognize the presence of a virus in a single cell. As a result, the behavior of the virus is studied at a population level, and you lose all the details you need—such as how the behavior of a single cell correlates to the number of copies of the virus that are penetrating the cell.
Once we understand how the virus enters a single cell and under what conditions, we will eventually be able to detect mutations. Though we will not be able to pinpoint which mutations will occur, we will be able to quantify how many base mutations there are, for instance. This information would help virologists assess how many mutations have to occur before the virus does not respond to a vaccine.
In essence, we are trying to understand how mutations affect the ecology of a virus. With this understanding, we can develop synthetic vaccines that alter this ecology. Think of a population of bees: If we disrupt the bee colony (ecology) by injecting a synthetic vaccine that masks itself as a worker bee by looking like a virus but lacking the infectious parts, it could be possible to disrupt the mechanisms of interaction to the point where the queen bee or entire colony ceases to function.
We are one year into the four-year project. If the tags are successful in detecting and monitoring the flu at the single-cell level, they could eventually be applied to other viruses.
One of the major goals in cancer research today is to reduce the toxicity of traditional therapies by eliminating cancerous cells while preserving healthy tissue. How do you synthesize and functionalize the SERS tags such that they not only go where they need to inside the human body but also survive the harsh conditions?
I teach a class on this very topic. Delivery is currently very inefficient. Only three to five percent of nanoparticles injected into the bloodstream of an animal reach the position they are supposed to. The kidneys, liver, and spleen clear the rest. My duty as a nanomaterials chemist is to design particles that will resist this circulation and exist for 24 to 48 hours without undergoing changes in their properties.
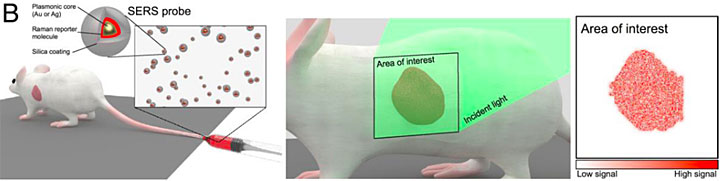
For in vivo analysis in mouse models, the SERS tags are injected in the animal via the tail vein and their bio-distribution and specific tissue targeting are followed using a microscope. Source: Memorial Sloan Kettering Cancer Center.
Here is where the discipline of bioconjugate chemistry comes into play—how can we make particles more resistant to enzymatic breakdown and more effective at reaching specific tissues and delivering drugs to those tissues?
A few years ago, my group had some initial promising results with gold nanoparticles. We found that when they absorb electromagnetic radiation, they not only increase the field around them but also heat up substantially. Tumor tissue needs to reach 32 to 47 degrees Celsius [89.6 to 116.6 degrees Fahrenheit] to die. So if these gold nanoparticles are able to reach the cancerous tissue, the heat that is concentrated around the particles will propagate to the tissue, killing the cancer cells. Even more interesting is that when we bind a molecule of doxorubicin—a chemotherapy drug that slows or stops the growth of cancer cells—to these gold nanoparticles, bring the molecule-nanoparticle conjugate to the tissue, and apply heat, cancer cells are killed much more effectively. There is a cooperative effect between the two therapies.
This method (technically known as photothermal therapy) is the only FDA-approved one with metal nanoparticles. So far, it is being used to target skin cancer. The near-infrared light used in photothermal therapy cannot deeply penetrate beyond the skin’s surface into tissues because it gets absorbed and scattered (though to a lesser extent than visible light, which is readily absorbed by water in the body). My group is playing with this wavelength range (800 nanometers and up). When making particles, we need to make sure they absorb radiation in that window. That way, when we deliver the particles to the tissue, the light is effectively absorbed by the nanoparticle tags to increase sensitivity. The nanostars I mentioned can be tuned to absorb this light very well.
A question I used to get at a lot of conferences regarding the gold nanostars was “Won’t the spikes poke the cell and the cell will die?” I told one of the students in my group that we needed to do a detailed study to put this question to rest. Our yearlong study addressed whether the surface chemistry (what molecules we put on the nanoparticles), shape, or size of the nanoparticles is toxic to cells, and if the particles are more toxic to cancerous or healthy cells. We also investigated mechanisms of toxicity.
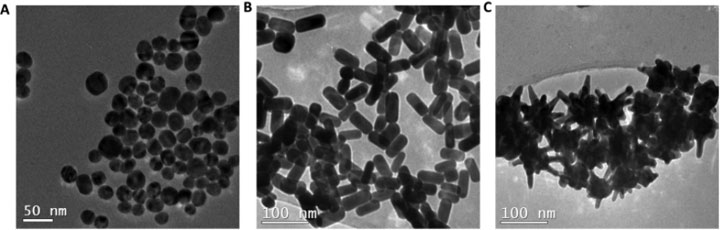

Transmission electron microscope images of (A) nanospheres, (B) nanorods, and (C) nanostars coated with a compound known as CTAB. CTAB-coated particles were found to be the most toxic to cells, causing membrane damage to both cancerous and healthy cells. Source: Bioconjugate Chemistry 2017, 28: 449–460.
My student ran 96 cell cultures in parallel with human glioblastoma and human dermal fibroblast cell lines, and we found that the composition (surface chemistry) determines toxicity. Compared to nanospheres and nanorods, nanostars coated with PEG [poly(ethylene glycol)] were the least toxic and most biocompatible. Without the PEG coating, the particles clump up. PEG creates a buffer that makes the cell believe that the particle is ok for the body—kind of like a camouflage because the cell does not recognize the particle. Upon exposure to the nanoparticles, healthy cells die through necrosis and cancer cells through forced apoptosis (cell suicide). We observed higher viabilities in the PEG-coated gold nanoparticles. This result may be because most of the nanoparticles are taken up by the cells in membrane-bound vesicles, limiting their interaction with organelles and the nucleus and thus reducing toxicity.
How would the SERS tags detect a cancer like leukemia, in which cancer cells circulate throughout the bloodstream and are not localized in a tumor mass?
We have to figure out how to concentrate those cells circulating in the bloodstream. Circulating tumor cells are an important problem in oncology. My group recently figured out a way to enrich samples of cells with aptamer-functionalized platforms.
Aptamers are DNA or RNA strands that are designed to work just like antibodies. In the presence of a specific target, they fold in a particular conformation and electrostatically bind the target molecules. For instance, proteins have epitopes—pockets with a high charge distribution at the surface—and when aptamers fold, they fit into the pockets of these proteins like a lock and key. The idea is to flow cells over a glass microscope slide coated with aptamers that have an affinity for cancer biomarkers. Any cells containing those biomarkers should then stick to the slide. Currently, this process is not very effective; we expect to capture only 50 to 60 percent of those cells.
Have you investigated how the nanostars or differently shaped nanoparticles could be applied to other applications beyond the biomedical ones?
While I was doing my postdoc at UC Santa Barbara, the research group of Professor Bazan, for whom I was working for, focused on organic solar cells. This group demonstrated that if you add certain molecules to a polymer mixture, you modify the conformation and create preferential alignment of the polymer chain such that you improve charge carrier migration to two electrodes. The thought that elongated nanoparticles might be able to do the same always tickled me. When I became an independent investigator, I started collaborating with scientists at the Air Force Research Laboratory at Wright-Patterson Air Force Base in Ohio to understand the synthesis and growth of gold nanorods, and presented an idea: if we have elongated gold nanoparticles like nanorods, can we induce morphology changes in the polymer layer to create preferential pathways in solar cells? My hypothesis was that the plasmonic effect would not be as relevant as the morphology effect in gold nanoparticles. In testing this hypothesis, I found that morphology alone enhanced the efficiency of the solar cells by 30 percent because of the alignment.
Recently, my group started studying how nanostars can be used in catalysis. When you excite these nanostars with near-infrared radiation, you generate hot electrons at the spikes. Photocatalytic experiments have shown that in nanostars capped with the catalyst titanium dioxide, the location of the hot spots at the tip of the spikes enables the transfer of hot electrons through the metal-semiconductor barrier, improving photoreduction reaction rates.
To confidently use these nanostars in catalysis, sensing, or imaging, we need to be able to estimate the fundamental physical and optical properties of the particles with different diameters, spike lengths and densities, and material compositions. In 2017, I published a paper in collaboration with other Rutgers researchers and CFN electron microscopist Huolin Xin on this topic. In particular, we used high-resolution transmission electron microscopes at the CFN to produce three-dimensional images of the structure of the nanostars.
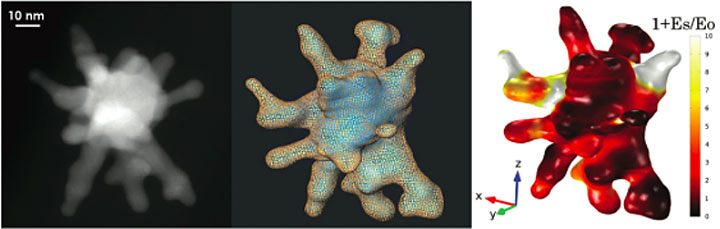
A three-dimensional topogram of the nanostar as captured with a transmission electron microscope (left), nanostar topography reconstructed with 22,000 tetrahedral elements (center), and scattered electric field distribution (right) calculated using modeling software. Source: Nanoscale, 2017, 9, 3766.
We transferred the resulting data into modeling software to understand the electric field properties and predict important properties—such as volume and surface area—of the nanostars. I am very excited about this work because our method could be used to understand the structure-property relationships of nanoparticles of any shape and chemical composition. Several scientists have contacted me to thank me for the method, and I am so happy to know that I have helped others move along in their work.
What equipment do you use at the CFN, and what is it like being a CFN user?
We use fluorometers to determine the presence and concentration of molecules in our samples and transmission electron microscopes to image our nanoparticles and understand how to optimize their morphology for cellular localization.
I have been a user at the CFN since I started at Rutgers in 2009, and coming to the CFN was one of the best decisions I have made. I had known about Brookhaven Lab prior to that time, but my colleagues at Rutgers told me that my research seemed to be spot on for the capabilities at the CFN. At the CFN, you not only get access to top-notch facilities but also to scientists who help you with your projects or collaborate with you. Now, I recommend the CFN to every new professor to Rutgers.
Personalized or precision medicine is a buzzword nowadays. Do you see the SERS tags becoming part of personalized cancer detection and treatment?
First, we need to do a lot more work in understanding the genome of an individual person. Then a comparison of the genomes between family members could reveal patterns or mutations correlated with a specific disease. Even if the mutations are present in low concentration, the idea is that the SERS tags would be able to pick them up.
My group is now gearing up to detect specific biomarkers in blood. Another buzzword today is exosome: a vesicle secreted by most types of cells, including cancer cells, to remove waste. Exosomes contain information about the cell itself because they have the same membrane, and they are quite concentrated in the blood. Some proteins are packaged into exosomes. From a personalized medicine standpoint, exosomes could serve as diagnostic tools and biomarkers for disease.
How did you get into science and extend yourself to so many different areas?
I was never one of those kids who said I want to be a scientist when I grow up. I think I wanted to be a scientist all along but just did not know it. I have always been very curious about how things work, and when I was little, I begged my mom for a microscope but she never bought me one. At my science high school that I attended in Italy, I got interested in chemistry. I remember there was a chemistry Olympiad, and my teacher told me that he did not take girls for the competition but he would allow me to go. I ended up doing better than anyone else!
I studied physical chemistry as an undergraduate at the University of Padova, where I pursued my PhD in chemical sciences. Then I wanted to see how research works in the United States. My plan was to stay for a year to do a postdoc, but here I am 12 years later!
All of the things that I do now are what I like to do. I am a physical chemist by training but I have incorporated my early interest in biology into my research. The SERS tags are very much physics and biology, and part of the reason that I started investigating the use of SERS for early cancer detection is that my grandfather died of brain cancer. I have been working on SERS in the context of cancer for the past decade. I guess you could say I have reached the thermodynamic minimum of my passions!
Nanoparticles have always been at the core of my research, but I like to bring my expertise to different areas and challenge myself. I am always looking for how I can apply the SERS technique to other fields and enjoy being challenged by problems outside my area of expertise.
Some of my research is also the result of serendipity. For example, the reason I got involved with the virology project at DARPA is because I had initially contacted a program manager about using the tags to detect small differences in the production of proteins in people who experience disruptions in their circadian rhythms—such as soldiers who are awake during the night. The manager told me that program was ending but there was a new program in virology. I went to proposal day and ended up finding four other investigators who became part of the team on the virology project that DARPA funded. This project represents the first time that SERS will be used in virology. I constantly seek opportunities to put myself out there because I want to show the scientific community that the technique can be applied to solve a broad range of problems.
Brookhaven National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
Follow @BrookhavenLab on Twitter or find us on Facebook.
2018-12892 | INT/EXT | Newsroom