Accelerated Computing Hackathon Returns for Second Year
Participants in the five-day coding workshop at Brookhaven Lab harnessed the processing power of NVIDIA graphics processing units to reduce time to discovery in astrophysics, particle physics, biology, and other scientific applications
December 14, 2018
 enlarge
enlarge
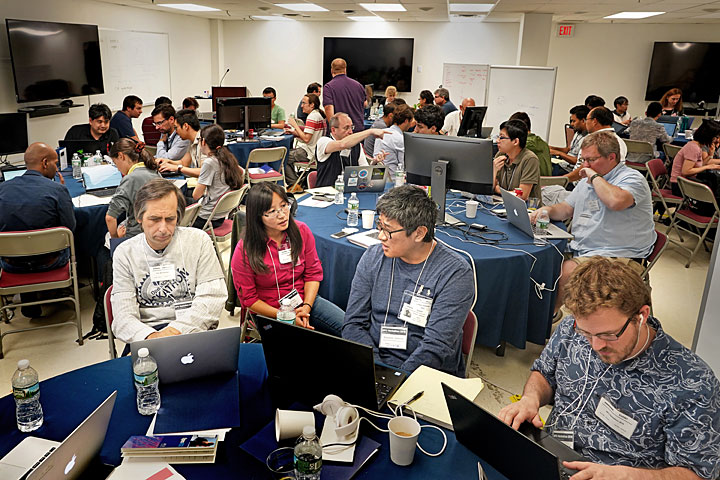
From Sept. 17 through 21, teams representing various universities, national labs, and technology companies came to Brookhaven Lab's second graphics processing unit (GPU) hackathon—hosted by the Computational Science Initiative—to accelerate their scientific application codes.
From simulating the universe’s formation to developing machine learning-based technology of the future, modern scientific discoveries increasingly depend on our ability to rapidly perform computations on an extremely large amount of data. Graphics processing units (GPUs), or massively parallel computing devices with high memory bandwidth, are designed to handle such big data by using thousands of computing cores that operate together. In order for scientific applications to take advantage of this accelerated computing, they must be ported to and optimized on GPUs. However, because the GPU architecture is significantly different from that of traditional central processing units (CPUs), different programming techniques are required.
A learning model
In June 2017, the Computational Science Initiative (CSI) at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory hosted its first GPU hackathon, called “Brookathon,” to help bridge this learning gap. For five days, GPU programming experts from national labs, universities, and technology companies trained 10 teams of CPU-GPU system users on GPU programming techniques. Each team came with a different scientific application to accelerate on GPUs. The event was part of the Oak Ridge Leadership Computing Facility (OLCF)—a DOE Office of Science User Facility—GPU Hackathon series.
“Last year’s GPU hackathon was quite successful, with most teams achieving application performance improvements on GPUs,” said hackathon coordinator and co-organizer Meifeng Lin, a CSI computational scientist. “Given this success and the growing demand for using GPUs in scientific computing, CSI decided to offer the hackathon again this year.”
The event, held from Sept. 17–21, was one of seven hackathons in the 2018 OLCF GPU Hackathon series. It was jointly organized with DOE’s Oak Ridge National Laboratory (ORNL), Stony Brook University (SBU), the University of Delaware, and—new for this year—Fluid Numerics, LLC.
“Joseph Schoonover from Fluid Numerics was a mentor at last year’s Brookathon,” said Lin. “This year, he returned to help moderate the hackathon and provide some GPU resources for the participants to test their codes on the Google Cloud Platform.”
A video providing an overview of the hackathon and featuring interviews with mentors and participants.
“It is really nice to stay at the cutting edge of the software and hardware that is available,” said Schoonover. “Every time that I’ve come to a hackathon, I’ve learned a lot about where this industry is headed and what we can do to catch people up.”
Diverse scientific applications
Of the 13 team applications received, 10—seven new and three returning teams—were accepted. The teams came with applications written in a variety of programming languages, including Python, C, and Fortran. Mentors representing more than 20 GPU experts from Brookhaven Lab, Grinnell College, Mentor Graphics, NVIDIA, SBU, University of Delaware, and University of Tennessee at Knoxville helped the teams apply different programming models (e.g., CUDA, OpenACC, CuPy) to accelerate their codes. Throughout the week, the teams used performance profiling tools to analyze bottlenecks and track improvements. The teams’ scientific applications spanned diverse fields, including astrophysics, high-energy physics, nuclear physics, machine learning, aerospace, biology, chemistry, and materials science.
For example, a team representing General Electric (GE) and the University of Kansas came to the hackathon with a code for simulating fluid flow under the kind of conditions that would be experienced in a gas turbine engine. Their goal was to port the large code (roughly 100,000 lines) from CPU to GPU.
 enlarge
enlarge
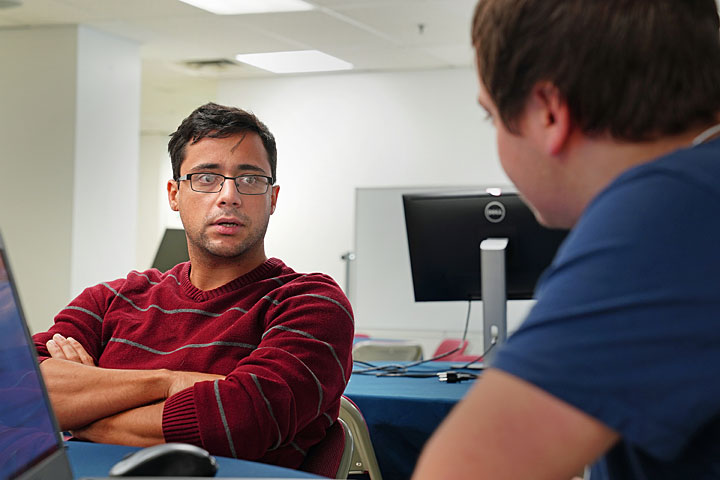
At the hackathon, participant Carlos Velez (left) of GE Global Research and mentor Kyle Friedline of the University of Delaware work on porting a computational fluid dynamics code from CPU to GPU.
“With this code, we can predict the performance of a new design for a machine without ever having to build it,” said team member Carlos Velez, a lead combustion modeling engineer at GE Global Research. “If we can make the code run faster—say, by halving the computing time—that equates to double the design iterations. The more design changes we explore, the better chance we have of reaching the best-performing design.”
By the end of the week, the team successfully ported some of their code to GPU. They will continue to port the hybrid code to a purely GPU version, and optimize it for parallel computational fluid dynamics simulations on leadership-class computing systems.
Team Xbiohack from the Simmerling Group at nearby SBU came to the hackathon to port one of the codes of Amber, a molecular dynamics software package, from CPU to GPU. Using Amber, scientists can simulate how biomolecules such as proteins and nucleic acids move and interact over time, atom by atom.
 enlarge
enlarge
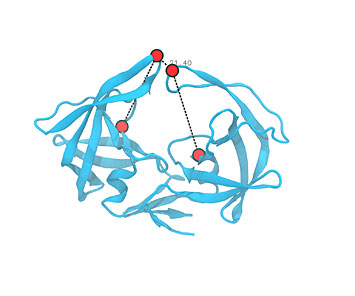
A molecular dynamics simulation of human immunodeficiency virus (HIV) protease, a protein that is vital to the HIV lifecycle. The Amber code restrains particular atoms of interest by grouping them into four points defining a dihedral. HIV protease was one of Team Xbiohack's sources of inspiration for their project at the hackathon.
“We use this code to restrain particular sets of atoms in particular ways,” explained team member and graduate student Zachary Fallon.
“Simulations with restrained atoms are relevant to several applications, including protein modeling and drug design.”
“Labs all over the world use Amber,” said team member and graduate student Lauren Raguette. “Porting this code to GPUs and optimizing it not only will benefit research in our own lab but also that of other members of the Amber community.”
“Over the last several years, the Amber community has been asking for this code to run on GPUs,” added team member and graduate student Kellon Belfon. “A simulation that take three weeks on CPUs can be completed within two days on GPUs.”
According to Raguette, the team achieved a 10-fold speedup over the original CPU-based code, and they will continue to optimize kernels—the computationally intensive parts of the calculation—back at SBU.
 enlarge
enlarge
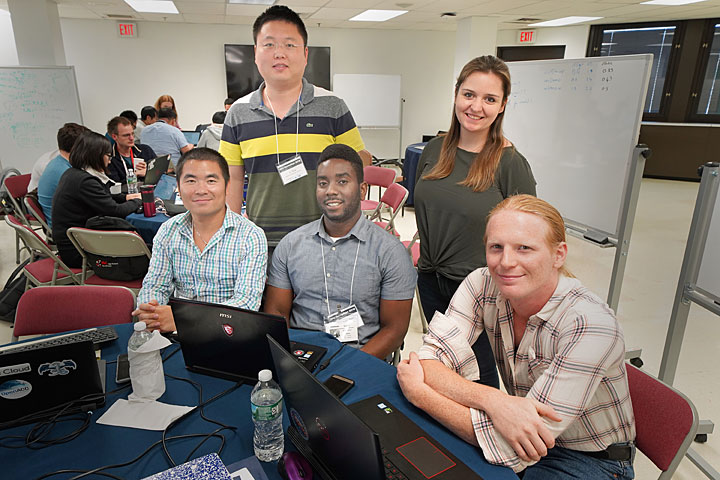
Team Xbiohack—(left to right) Daniel Tian (NVIDIA), Li Tang (Brookhaven Lab), Kellon Belfon (SBU), Lauren Raguette (SBU), and Zach Fallon (SBU)—ported a code to GPU that restrains certain parts of biomolecules, such as DNA and proteins.
“Not only were we able to improve our code during the week but also we gained knowledge and skillsets that we are already sharing with our fellow lab members,” said Raguette.
Another team focused their efforts on codes that are part of an astrophysics simulation suite.
“For one of the codes, we worked on incorporating additional physics capabilities to run on GPUs so that we can study more real-world problems,” said Maria (Lupe) Barrios Sazo, a PhD student in SBU’s Physics and Astronomy Department. “We also started from scratch to port another code for simulating the explosion of stars into supernovae.”
“When the week started, we had one physics solver working on GPU, but the code was not ported enough to run any scientifically meaningful simulation,” said mentor Max Katz, a solutions architect at NVIDIA. “By the end of the week, we were able to run one of our real science problems—one that includes nuclear reactions, which are needed to study supernovae—on GPUs. Because of this accomplishment, we are much better prepared to run the latest generation of GPU-powered supercomputers, such as Summit at Oak Ridge National Laboratory.”
 enlarge
enlarge
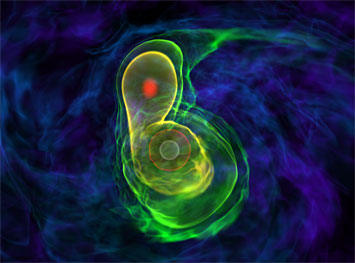
A simulation of two white dwarfs merging, one of the real-world problems that Barrios Sazo and the rest of Team AstroGPU worked on. It is possible that Type Ia supernovae—bright, brief pulses of light in the distant universe—originate from this explosive merger. Source: Katz et al. 2016.
Kazuhiro Terao, a physicist at DOE’s SLAC National Accelerator Lab, and his team had similar success in optimizing their machine learning (ML) algorithm for reconstructing and analyzing data from particle detectors called liquid-argon time-projection chambers. Physicists need very high-resolution images to identify which particles are present and determine how they are interacting with one another.
“The ML techniques we employ involve lots of massively parallel linear algebra operations, which can benefit from GPU hardware acceleration,” said Terao. “We have been using GPUs for our ML projects through high-level software frameworks provided by Google, Facebook, and other tech companies. But as we enhance the sophistication of our algorithm, these frameworks begin to suffer from long compute time.”
Within the first few days of debugging at the hackathon, Terao’s team found a few bottleneck operations, which they replaced with their own custom operations. This change decreased their compute time by more than a factor of three. The team also implemented a custom GPU code to reduce the frequency of CPU-GPU data transfer during the algorithm training process and split a huge ML algorithm on multiple GPUs.
 enlarge
enlarge
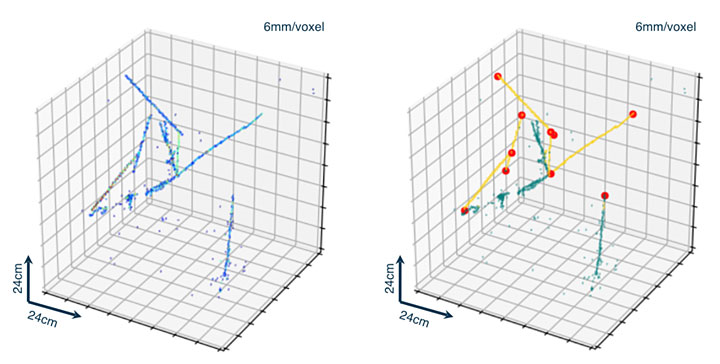
Left: input image showing charged particle trajectories in a detector. Right: annotated image using machine learning algorithms, where trajectories are classified into two different colors based on geometrical features, and edges of particle trajectories (red circles) are detected.
“As we try to solve more complex problems, our ML algorithm uses more memory, eventually to a level where it cannot fit on a single GPU,” explained Terao. “Establishing a generic technique to construct an ML algorithm on multiple GPUs would address this issue and would be helpful beyond our application.”
By the end of the hackathon, Terao’s team implemented a data parallelism scheme in which a single algorithm instance spans two GPUs.
Team SBNbuggy from Nevis Laboratories—a high-energy physics research center at Columbia University—also explored the use of GPUs for SBNfit. Physicists use this software framework to fit multiple measurements across different neutrino (a kind of subatomic particle) detectors and experiments. SBNfit is currently being used for the MicroBooNE neutrino experiment at DOE’s Fermilab, which will investigate neutrino properties and interactions. During the hackathon, the team focused on enabling GPU acceleration for two core computational procedures in SBNfit, achieving speedups as high as 200 times faster than the CPU-based version.
“It is gratifying to see how much progress these teams can make in such a short time with the hands-on guidance of their mentors,” said Lin. “With the increasing volume and velocity of data coming from various experimental facilities and theory-based simulations, CSI is committed to providing more training events to the scientific community to help them harness modern computational resources to advance science.”
A third GPU hackathon will be hosted at Brookhaven Lab next year from Sept. 23–27.
The hackathon received support from DOE’s High Energy Physics Center for Computational Excellence and Exascale Computing Project (ECP), and was sponsored by Fluid Numerics, Google, NVIDIA, and OpenACC, and ORNL.
Brookhaven National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
Follow @BrookhavenLab on Twitter or find us on Facebook.
2018-13093 | INT/EXT | Newsroom










