Scientists Decipher Structure of Nature’s ‘Light Switch’
New findings will help scientists understand how plants respond to light
May 31, 2010
UPTON, NY — When the first warm rays of springtime sunshine trigger a burst of new plant growth, it’s almost as if someone flicked a switch to turn on the greenery and unleash a floral profusion of color. Opening a window into this process, scientists at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory and collaborators at the University of Wisconsin, Madison, have deciphered the structure of a molecular “switch” much like the one plants use to sense light. Their findings, described online in the Proceedings of the National Academy of Sciences the week of May 31, 2010, help explain how the switch works and could be used to design new ways to modify plant growth.
Previous studies showed that the light-sensing structure, called a phytochrome, exists in two stable states. Each state is sensitive to a slightly different wavelength, or color, of light — from red to “far red,” which is close to the invisible infrared end of the light spectrum. As the phytochrome absorbs photons of one wavelength or the other, it changes shape and sends signals that help plants know when to flower, produce chlorophyll, and grow.
“The phytochrome is almost like nature’s light switch,” said Brookhaven biophysicist Huilin Li, who is also an associate professor at Stony Brook University and a lead author on the study. “Finding out how this switch is flipped on or off by a signal as subtle as a single photon of light is fascinating.”
 enlarge
enlarge
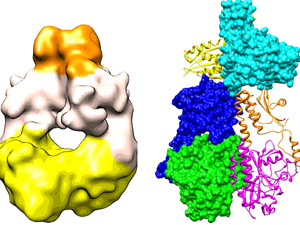
Left: The newly derived 3D map of a bacterial phytochrome dimer, produced using cryo electron microscopy. Right: By fitting x-ray crystal structures of several homologous fragments into this map, scientists have created an atomic model of the whole structure. The two monomers making up the complete structure — one shown as a “ribbon” diagram, the other using a space-filling display — dimerize in parallel with the two polypeptides intimately twisting around each other.
As with all biological molecules, one key to the phytochrome’s function is its structure. But scientists trying to get a molecular-level picture of a phytochrome have a formidable challenge: The phytochrome molecule is too dynamic to capture in a single image using techniques like x-ray crystallography. So, scientists have studied only the rigid and smaller pieces of the molecule, yielding detailed, but fragmented, information.
Now using additional imaging and computational techniques, the Brookhaven researchers and their collaborators have pieced together for the first time a detailed structure of a whole phytochrome.
Li and his collaborators studied a phytochrome from a common bacterium that is quite similar in biochemistry and function to those found in plants, but easier to isolate. Plant biologist Richard Vierstra of the University of Wisconsin provided the purified samples.
At Brookhaven, Li’s group used two imaging techniques. First, they applied a layer of heavy metal dye to the purified phytochrome molecules to make them more visible, and viewed them using an electron microscope. This produced many two-dimensional images from a variety of angles to give the researchers a rough outline of the phytochrome map.
The scientists also froze the molecules in solution to produce another set of images that would be free of artifacts from the staining technique. For this set of images, the scientists used a cryo-electron microscope.
Using computers to average the data from each technique and then combine the information, the scientists were able to construct a three-dimensional map of the full phytochrome structure. The scientists then fitted the previously determined detailed structures of phytochrome fragments into their newly derived 3-D map to build an atomic model for the whole phytochrome.
Though the scientists knew the phytochrome was composed of two “sister” units, forming a dimer, the new structure revealed a surprisingly long twisted area of contact between the two individual units, with a good deal of flexibility at the untwisted ends. The structure supports the idea that the absorption of light somehow adjusts the strength or orientation of the contact, and through a series of conformation changes, transmits a signal down the length of the molecular interface. The scientists confirmed the proposed structural changes during photo-conversion by mutagenesis and biochemical assay.
The scientists studied only the form of the phytochrome that is sensitive to red light. Next they plan to see how the structure changes after it absorbs red light to become sensitive to “far red” light. Comparing the two structures will help the scientists test their model of how the molecule changes shape to send signals in response to light.
This research was supported by Brookhaven’s Laboratory Directed Research and Development program, the National Institutes of Health, the National Science Foundation, and a grant from the University of Wisconsin College of Agricultural and Life Science.
2010-11143 | INT/EXT | Newsroom