Scientists Produce 3-D Chemical Maps of Single Bacteria
Researchers at NSLS-II used ultrabright x-rays to generate 3-D nanoscale maps of a single bacteria's chemical composition with unparalleled spatial resolution
November 16, 2018
 enlarge
enlarge
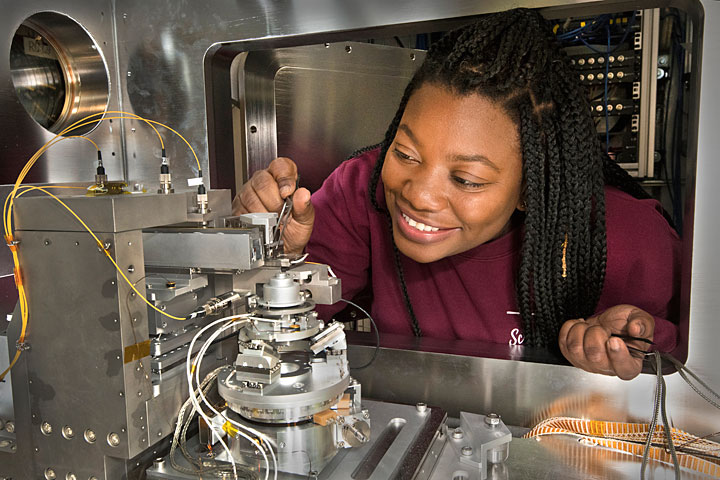
NSLS-II scientist Tiffany Victor is shown at the Hard X-ray Nanoprobe, where her team produced 3-D chemical maps of single bacteria with nanoscale resolution.
Scientists at the National Synchrotron Light Source II (NSLS-II)—a U.S. Department of Energy (DOE) Office of Science User Facility at DOE’s Brookhaven National Laboratory—have used ultrabright x-rays to image single bacteria with higher spatial resolution than ever before. Their work, published in Scientific Reports, demonstrates an x-ray imaging technique, called x-ray fluorescence microscopy (XRF), as an effective approach to produce 3-D images of small biological samples.
“For the very first time, we used nanoscale XRF to image bacteria down to the resolution of a cell membrane,” said Lisa Miller, a scientist at NSLS-II and a co-author of the paper. “Imaging cells at the level of the membrane is critical for understanding the cell’s role in various diseases and developing advanced medical treatments.”
The record-breaking resolution of the x-ray images was made possible by the advanced capabilities of the Hard X-ray Nanoprobe (HXN) beamline, an experimental station at NSLS-II with novel nanofocusing optics and exceptional stability.
“HXN is the first XRF beamline to generate a 3-D image with this kind of resolution,” Miller said.
While other imaging techniques, such as electron microscopy, can image the structure of a cell membrane with very high resolution, these techniques are unable to provide chemical information on the cell. At HXN, the researchers could produce 3-D chemical maps of their samples, identifying where trace elements are found throughout the cell.
 enlarge
enlarge
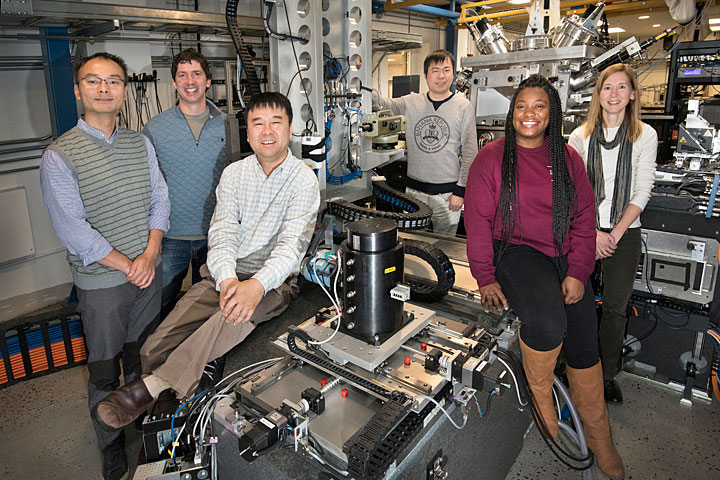
The research team at NSLS-II's Hard X-ray Nanoprobe. Pictured from left to right are Xiaojing Huang, Randy Smith, Yong Chu, Hanfei Yan, Tiffany Victor, and Lisa Miller.
“At HXN, we take an image of a sample at one angle, rotate the sample to the next angle, take another image, and so on,” said Tiffany Victor, lead author of the study and a scientist at NSLS-II. “Each image shows the chemical profile of the sample at that orientation. Then, we can merge those profiles together to create a 3-D image.”
Miller added, “Obtaining an XRF 3-D image is like comparing a regular x-ray you can get at the doctor’s office to a CT scan.”
The images produced by HXN revealed that two trace elements, calcium and zinc, had unique spatial distributions in the bacterial cell.
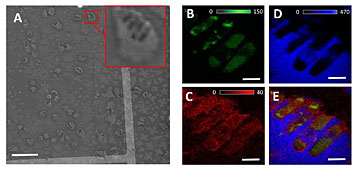
XRF images show the zinc (B), calcium (C), chlorine (D) distributions in the single bacteria. XRF image E shows all three elements in the cell. Image A shows bacteria embedded in sodium chloride crystals.
“We believe the zinc is associated with the ribosomes in the bacteria,” Victor said. “Bacteria don’t have a lot of cellular organelles, unlike a eukaryotic (complex) cell that has mitochondria, a nucleus, and many other organelles. So, it’s not the most exciting sample to image, but it’s a nice model system that demonstrates the imaging technique superbly.”
Yong Chu, who is the lead beamline scientist at HXN, says the imaging technique is also applicable to many other areas of research.
“This 3-D chemical imaging or fluorescence nanotomography technique is gaining popularity in other scientific fields,” Chu said. “For example, we can visualize how the internal structure of a battery is transforming while it is being charged and discharged.”
In addition to breaking the technical barriers on x-ray imaging resolution with this technique, the researchers developed a new method for imaging the bacteria at room temperature during the x-ray measurements.
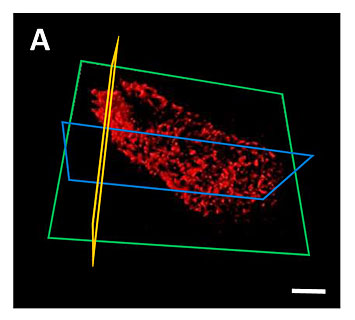
A 3-D view of the single bacteria produced through XRF.
“Ideally, XRF imaging should be performed on frozen biological samples that are cryo-preserved to prevent radiation damage and to obtain a more physiologically relevant understanding of cellular processes,” Victor said. “Because of the space constraints in HXN’s sample chamber, we weren’t able to study the sample using a cryostage. Instead, we embedded the cells in small sodium chloride crystals and imaged the cells at room temperature. The sodium chloride crystals maintained the rod-like shape of the cells, and they made the cells easier to locate, reducing the run time of our experiments.”
The researchers say that demonstrating the efficacy of the x-ray imaging technique, as well as the sample preparation method, was the first step in a larger project to image trace elements in other biological cells at the nanoscale. The team is particularly interested in copper’s role in neuron death in Alzheimer’s disease.
“Trace elements like iron, copper, and zinc are nutritionally essential, but they can also play a role in disease,” Miller said. “We’re seeking to understand the subcellular location and function of metal-containing proteins in the disease process to help develop effective therapies.”
The work was supported by DOE’s Office of Science, the National Institutes of Health, and the National Science Foundation.
Brookhaven National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
Follow @BrookhavenLab on Twitter or find us on Facebook.
2018-13105 | INT/EXT | Newsroom









