Understanding an Essential Chaperone Complex
Scientists reveal a protein binding mechanism that is connected to embryo development and cancer
September 30, 2020
 enlarge
enlarge
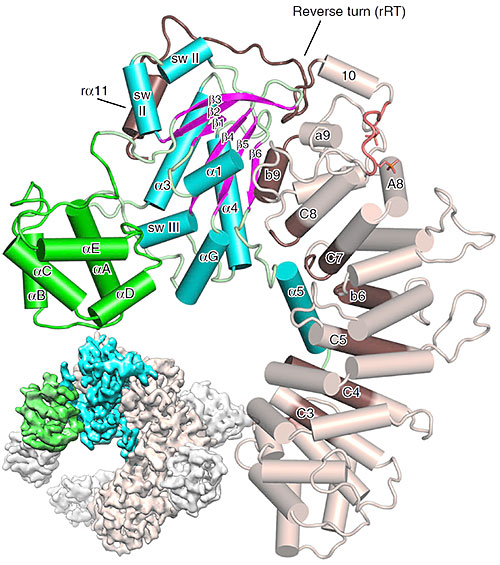
The small figure (bottom left) shows a schematic based on cryo-electron microscopy data, while the large figure in the forefront shows a detailed zoom in based on x-ray diffraction data. Image credit: L. J. McClelland, et. al. Nat. Comm. 11, 1077 (2020).
The Science
The structure of the chaperone Ric-8A bound to G protein alpha reveals the mechanism for Gα activation through phosphorylation of Ric-8A.
The Impact
Ric-8A is a protein involved in the regulation of cell division that is essential for embryo development. This structure reveals how it acts as a chaperone for Gα in this process.
Summary
Many of our modern developments in biology, medicine, and biotechnology have been made possible through our understanding of how biological structures such as proteins in cells interact with each other. But to truly reveal their function, as well as the role they play in diseases and medical conditions, scientists need to visualize these structures at the atomic level—especially for more elusive proteins and their functions, such as the Ric-8A protein, a Guanine Nucleotide exchange factor that serves as a chaperone for G protein alpha subunits (Gα). Gα are located at membranes and inside cells and are an essential part of several cellular signaling systems. Ric-8A is essential for asymmetric cell division in the process of embryonic development and is also a potential therapeutic target for certain cancers.
In this study, scientists revealed the structure and binding mechanism of the Ric-8A protein to the Gαi1 complex. The team has used a combination of cryo-electron microscopy and x-ray studies at various light source facilities. As part of their investigation, the team used x-ray crystallography at the Frontier Microfocusing Macromolecular Crystallography (FMX) beamline. The FMX beamline is part of the advance life science suite of beamlines at the National Synchrotron Light Source II (NSLS-II), a U.S. Department of Energy (DOE) Office of Science User Facility located at DOE’s Brookhaven National Laboratory.
Through their studies, the researchers discovered that the mechanism of Ric-8A differs from the usual mechanism that the G protein uses at its receptors. Ric-8A engages a specific conformation of Gα at multiple interfaces to form a complex that is stabilized by phosphorylation within a Ric-8A segment that connects two Gα binding sites.
Download the research summary slide
Contact
Stephen R. Sprang
University of Montana
Stephen.Sprang@mso.umt.edu
Publication
L. J. McClelland, K. Zhang, T.-C. Mou, J. Johnston, C. Yates-Hansen, S. Li, C. J. Thomas, T. I. Doukov, S. Triest, A. Wohlkonig, G. G. Tall, J. Steyaert, W. Chiu, S. R. Sprang. Structure of the G protein chaperone and guanine nucleotide exchange factor Ric-8A bound to Gαi1. Nature Communications 11, 1077 (2020). DOI: 10.1038/s41467-020-14943-4
Funding
This research was supported by NIH R01GM105993 (S.R.S.); NIH P41GM103832, R01GM079429, and S10OD021600 (W.C.); NSF 1738547 (T.-C.M); NIH R01- GM088242 (G.G.T.); resources of Instruct-ERIC, part of the European Strategy Forum on Research infrastructures (ESFRI), the Research Foundation-Flanders (FWO) for their support to Nanobody discovery, and the Strategic Research Program (SRP) of the Vrije Universiteit Brussel (S.T. and J.S.). The Integrated Structural Biology Core at the University of Montana Center for Biomolecular Structure and Dynamics is supported by NIH COBRE award P20GM103546 (S.R.S.). The FMX (17-ID-2) beamline is supported by NIH grant P41GM111244 and the Department of Energy (DOE), KP1605010. T.I.D. is supported by the SSRL Structural Molecular Biology Program by the DOE and by NIH grant P41GM103393. Small angle X-ray scattering experiments were conducted at the Advanced Photon Source, operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357 with support of NIH grant P41 GM103622 and 1S10OD018090-01 for purchase of the Pilatus 3 1M detector.
2020-18798 | INT/EXT | Newsroom









