New Computer Model Could Explain how Simple Molecules Took First Step Toward Life
Two Brookhaven researchers developed theoretical model to explain the origins of self-replicating molecules
July 28, 2015
 enlarge
enlarge
Brookhaven researchers Sergei Maslov (left) and Alexi Tkachenko developed a theoretical model to explain molecular self-replication.
UPTON, NY—Nearly four billion years ago, the earliest precursors of life on Earth emerged. First small, simple molecules, or monomers, banded together to form larger, more complex molecules, or polymers. Then those polymers developed a mechanism that allowed them to self-replicate and pass their structure on to future generations.
We wouldn't be here today if molecules had not made that fateful transition to self-replication. Yet despite the fact that biochemists have spent decades searching for the specific chemical process that can explain how simple molecules could make this leap, we still don't really understand how it happened.
"We now have this mechanism for the emergence of polymers that can potentially carry and transmit information, taking us from the primordial soup to a primordial soufflé."
— Brookhaven researcher Alexei Tkachenko
Now Sergei Maslov, a computational biologist at the U.S. Department of Energy's Brookhaven National Laboratory and adjunct professor at Stony Brook University, and Alexei Tkachenko, a scientist at Brookhaven's Center for Functional Nanomaterials (CFN), have taken a different, more conceptual approach. They've developed a model that explains how monomers could very rapidly make the jump to more complex polymers. And what their model points to could have intriguing implications for CFN's work in engineering artificial self-assembly at the nanoscale. Their work is published in the July 28, 2015 issue of The Journal of Chemical Physics.
To understand their work, let's consider the most famous organic polymer, and the carrier of life's genetic code: DNA. This polymer is composed of long chains of specific monomers called nucleotides, of which the four kinds are adenine, thymine, guanine, and cytosine (A, T, G, C). In a DNA double helix, each specific nucleotide pairs with another: A with T, and G with C. Because of this complementary pairing, it would be possible to put a complete piece of DNA back together even if just one of the two strands was intact.
While DNA has become the molecule of choice for encoding biological information, its close cousin RNA likely played this role at the dawn of life. This is known as the RNA world hypothesis, and it's the scenario that Maslov and Tkachenko considered in their work.
The single complete RNA strand is called a template strand, and the use of a template to piece together monomer fragments is what is known as template-assisted ligation. This concept is at the crux of their work. They asked whether that piecing together of complementary monomer chains into more complex polymers could occur not as the healing of a broken polymer, but rather as the formation of something new.
"Suppose we don't have any polymers at all, and we start with just monomers in a test tube," explained Tkachenko. "Will that mixture ever find its way to make those polymers? The answer is rather remarkable: Yes, it will! You would think there is some chicken-and-egg problem—that, in order to make polymers, you already need polymers there to provide the template for their formation. Turns out that you don't really."
Instilling memory
 enlarge
enlarge
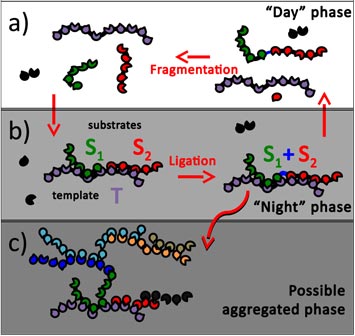
A schematic drawing of template-assisted ligation, shown in this model to give rise to autocatalytic systems.
Maslov and Tkachenko's model imagines some kind of regular cycle in which conditions change in a predictable fashion—say, the transition between night and day. Imagine a world in which complex polymers break apart during the day, then repair themselves at night. The presence of a template strand means that the polymer reassembles itself precisely as it was the night before. That self-replication process means the polymer can transmit information about itself from one generation to the next. That ability to pass information along is a fundamental property of life.
"The way our system replicates from one day cycle to the next is that it preserves a memory of what was there," said Maslov. "It's relatively easy to make lots of long polymers, but they will have no memory. The template provides the memory. Right now, we are solving the problem of how to get long polymer chains capable of memory transmission from one unit to another to select a small subset of polymers out of an astronomically large number of solutions."
According to Maslov and Tkachenko's model, a molecular system only needs a very tiny percentage of more complex molecules—even just dimers, or pairs of identical molecules joined together—to start merging into the longer chains that will eventually become self-replicating polymers. This neatly sidesteps one of the most vexing puzzles of the origins of life: Self-replicating chains likely need to be very specific sequences of at least 100 paired monomers, yet the odds of 100 such pairs randomly assembling themselves in just the right order is practically zero.
"If conditions are right, there is what we call a first-order transition, where you go from this soup of completely dispersed monomers to this new solution where you have these long chains appearing," said Tkachenko. "And we now have this mechanism for the emergence of these polymers that can potentially carry information and transmit it downstream. Once this threshold is passed, we expect monomers to be able to form polymers, taking us from the primordial soup to a primordial soufflé."
While the model's concept of template-assisted ligation does describe how DNA—as well as RNA—repairs itself, Maslov and Tkachenko's work doesn't require that either of those was the specific polymer for the origin of life.
"Our model could also describe a proto-RNA molecule. It could be something completely different," Maslov said.
Order from disorder
The fact that Maslov and Tkachenko's model doesn't require the presence of a specific molecule speaks to their more theoretical approach.
This video explores a new model, fully described in The Journal of Chemical Physics, from AIP Publishing, which proposes a potential mechanism for how self-replication could have emerged.
"It's a different mentality from what a biochemist would do," said Tkachenko. "A biochemist would be fixated on specific molecules. We, being ignorant physicists, tried to work our way from a general conceptual point of view, as there's a fundamental problem."
That fundamental problem is the second law of thermodynamics, which states that systems tend toward increasing disorder and lack of organization. The formation of long polymer chains from monomers is the precise opposite of that.
"How do you start with the regular laws of physics and get to these laws of biology which makes things run backward, which make things more complex, rather than less complex?" Tkachenko queried. "That's exactly the jump that we want to understand."
Applications in nanoscience
The work is an outgrowth of efforts at the Center for Functional Nanomaterials, a DOE Office of Science User Facility, to use DNA and other biomolecules to direct the self-assembly of nanoparticles into large, ordered arrays. While CFN doesn't typically focus on these kinds of primordial biological questions, Maslov and Tkachenko's modeling work could help CFN scientists engaged in cutting-edge nanoscience research to engineer even larger and more complex assemblies using nanostructured building blocks.
"There is a huge interest in making engineered self-assembled structures, so we were essentially thinking about two problems at once," said Tkachenko. "One is relevant to biologists, and second asks whether we can engineer a nanosystem that will do what our model does."
The next step will be to determine whether template-aided ligation can allow polymers to begin undergoing the evolutionary changes that characterize life as we know it. While this first round of research involved relatively modest computational resources, that next phase will require far more involved models and simulations.
Maslov and Tkachenko's work has solved the problem of how long polymer chains capable of information transmission from one generation to the next could emerge from the world of simple monomers. Now they are turning their attention to how such a system could naturally narrow itself down from exponentially many polymers to only a select few with desirable sequences.
"What we needed to show here was that this template-based ligation does result in a set of polymer chains, starting just from monomers," said Tkachenko. "So the next question we will be asking is whether, because of this template-based merger, we will be able to see specific sequences that will be more 'fit' than others. So this work sets the stage for the shift to the Darwinian phase."
This work was supported by the DOE Office of Science.
Brookhaven National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
2015-11745 | INT/EXT | Newsroom









