COVID-19 Research
Combining expertise across disciplines to address drug development, information processing, and more
Brookhaven National Laboratory has exceptional resources for addressing some of the most urgent scientific and logistical challenges of the COVID-19 pandemic.
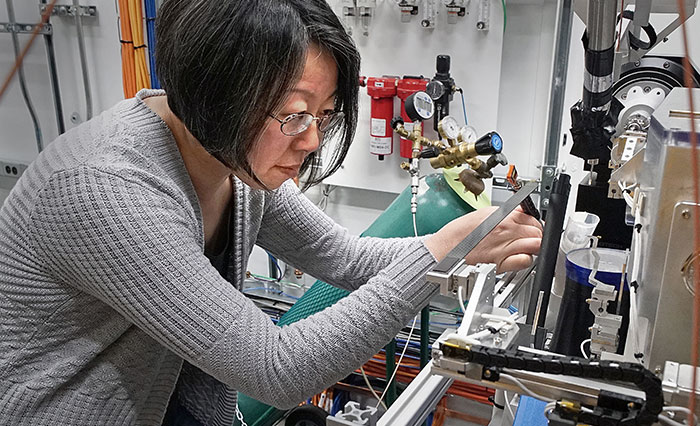

Structural biology efforts
Experiments running at two protein crystallography beamlines and one solution scattering beamline at the National Synchrotron Light Source II (NSLS-II) aim to characterize the atomic-level structure of viral components, including how they connect with receptors on human cells, so scientists can identify ways to block the infection-causing interactions. NSLS-II has the brightest synchrotron x-rays in the world for studying the proteins in the virus and understanding the processes hijacked in infected cells.
Additionally, researchers use the cryo-electron microscopes of the Laboratory for BioMolecular Structure (LBMS), a research facility located at Brookhaven Lab, for studying the various proteins of the virus. LBMS offers complementary imaging techniques to the efforts at NSLS-II.
Scientists from several pharmaceutical companies, academic researchers, and Brookhaven scientists are engaged in work at both facilities, often collaborating in the search for antiviral agents and targets for vaccines.
Researchers, submit a proposal for COVID-19-related research using the Rapid Access form.
See list of computation projects
Computational approaches
Narrowing the search for drugs
Once scientists know the viral components they want to target for developing therapeutic agents, they need to know which small drug-like molecules will fit into the viral-protein pockets or the cell-surface receptors to which the virus binds. Computational scientists at Brookhaven and collaborating institutions are helping to speed up the search by modeling proteins that play critical roles in the virus life cycle to identify promising drug targets. Using supercomputing resources across the U.S., the scientists will screen the chemical structures of a wide range of small molecules that might be able to block key viral functions. These “virtual” experiments will explore the chemical structures of known drugs that are already licensed and could quickly be repurposed, as well as libraries of small drug-like molecules that could be developed into drugs.
Tracking research efforts
The number of new papers related to COVID-19 that have been published since the start of the outbreak in December 2019 is rapidly growing. Brookhaven’s computational scientists are developing a “natural language processing” program that will search through a library of all these papers to pull out the most relevant information based on researchers’ plain-language questions. Using this system, scientists would be able to more easily find and track the latest data on which drugs are in clinical trials, for example, or pull out research on the latest potential targets.
Modeling disease spread
A computational model developed with help from Brookhaven scientists at the Center for Functional Nanomaterials was designed to simulate the spread of the virus in the Chicago area. Government officials in Illinois used this model in an early effort to optimize their response to the outbreak there, and ongoing research continues to improve and extend this work.
Special note to the research community: How can we help with your COVID-19 research needs? Let us know.
See AllRelated News

Brookhaven Lab Biophysicist F. William Studier Awarded Merkin Prize in Biomedical Technology
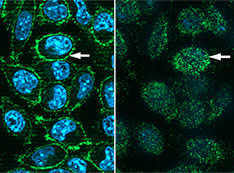
Scientists Make COVID Receptor Protein in Mouse Cells
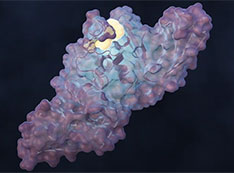
ORNL-led Team Designs Molecule to Disrupt SARS-CoV-2 Infection

Department of Energy Announces $5 Million for Research to Develop New Models for Bio-Preparedness
Press Coverage
-
Nobel Prize for mRNA Vaccines
-
Around The World, In 40 Years, On The T7 Express
-
High-precision construction of sPHENIX detector components wraps up at NPL
- Newsclips are articles in the press that are either about Brookhaven or are of interest to the Brookhaven community. Posting of these articles does not imply an endorsement of their content.




