Structure of a Nuclear Pore Complex's "Ticket System"
Scientists revealed the 3D structure of "gatekeeper" proteins for which failure is associated with many diseases
July 31, 2018
 enlarge
enlarge
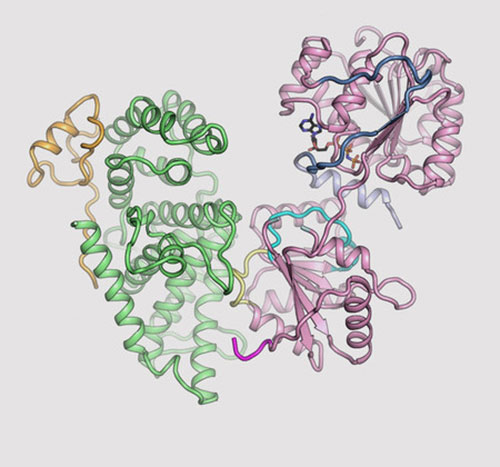
The image shows the three-dimensional (3D) structures for three proteins in the human nuclear pore complex: Gle1 (green), Nup42 (yellow), and DDX19 (pink). Image credit: Nature Commun. 9, 2319 (2018)
The Science
Scientists revealed the 3D structure of a nuclear pore complex’s “ticket system”, the protein complex Gle1-Nup42, from three organisms and showed how it ensures mRNA transport in humans.
The Impact
Nuclear pore complexes (NPCs) are the “gatekeepers” between a cell’s nucleus and its main chamber. Defects in transfer through NPCs are associated with many diseases.
Summary
Proteins are the main actors in many biological functions and their failure is often connected to various diseases. The nuclear pore complex (NPC) is a protein that acts as a “gatekeeper” between a cell’s nucleus and its main chamber. These special proteins control the passage of macromolecules between the double lipid membrane dividing the nucleus from the cytoplasm. Problematic transfer through this barrier is connected to several diseases. Even through this protein serves such an important function, we don’t actually know how the NPC directly participates in macromolecular transports.
One of the transports handled by NPCs is the transfer of messenger ribonucleic acid (mRNA) from the nucleus into the main chamber. mRNA is a set of instructions, which was copied from the deoxyribonucleic acid (DNA) of the cell, for the synthesis of proteins.
At the beginning of the transfer, a specific complex, called Nxf1•Nxt1, attaches itself to the mRNA string. This complex acts as “the ticket” for the mRNA to take a ride through the barrier. After the mRNA string passes through the barrier, the ticket needs to be removed so that the mRNA is not accidently send back. In the final step of mRNA transfer, a protein complex called nucleoporins Gle1, Nup214, and Nup42, activates a different protein, the helicase DDX19, that can remove the “ticket” from mRNA.
In this study, the scientists crystalized the protein complex Gle1•Nup42 from different organisms, including humans. The team used x-rays from the Frontier Microfocusing Macromoledular Crystallography (FMX) beamline to reveal the 3-D crystal structure of the protein complex for all three different organisms. The FMX beamline is part of the life science research suite at the National Synchrotron Light Source II (NSLS-II), a U.S. Department of Energy (DOE) Office of Science User Facility located at DOE’s Brookhaven National Laboratory.
The scientists were able to uncover the evolutionarily conserved binding mode for each organism. They learned that while fungi require an additional step during activation, this step is not required in humans.
Their results provide the foundation for further mechanistic analyses of mRNA export in humans and its connection to diseases.
Contact
André Hoelz
California Institute of Technology
hoelz@caltech.edu
Publication
D. H. Lin, A. R. Correia, S. W. Cai, F. M. Huber, C. A. Jette, A. Hoelz. Structural and functional analysis of mRNA export regulation by the nuclear pore complex. Nature Commun. 9, 2319 (2018). DOI: 10.1038/s41467-018-04459-3
Funding
This work was performed at the Frontier Microfocusing Macromolecular Crystallography (FMX) beamline at the National Synchrotron Light Source II (NSLS-II), SSRL Beamline 12-2, and the National Institute of General Medical Sciences and National Cancer Institute Structural Biology Facility (GM/CA) at the Advanced Photon Source (APS). This research used resources of the National Synchrotron Light Source II, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Brookhaven National Laboratory under Contract No. DE-SC0012704. SSRL and APS are also supported by the US Department of Energy. We acknowledge the Gordon and Betty Moore Foundation, the Beckman Institute, and the Sanofi-Aventis Bioengineering Research Program for their support of the Molecular Observatory at the California Institute of Technology. GM/CA has been funded in whole or in part with federal funds from the National Cancer Institute (grant ACB-12002) and the National Institute of General Medical Sciences (grant AGM-12006). The crystal structure of ctGle1CTD•Nup42CTD•IP6 was determined at the CCP4/APS School for Macromolecular Crystallography (2016), which was supported by the National Cancer Institute (AGM-12006) and the National Institute of General Medical Science (ACB-12002). D.H.L. and C.A.J. were supported by an NIH Research Service Award (5T32 GM07616). D.H.L. and F.M.H. were supported by an Amgen Graduate Fellowship through the Caltech-Amgen Research Collaboration. Scientists were also supported by NIH grant R01-GM117360.
2018-18955 | INT/EXT | Newsroom









