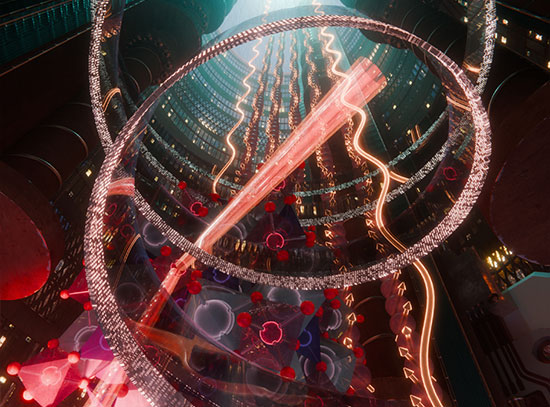3D Nanoscale Imaging of Proteins in Cells
Scientists used lanthanide-binding tags to image proteins at the level of a cell membrane, opening new doors for studies on health and medicine
March 31, 2020
3-D x-ray movie of a single E.coli bacteria where yellow represents the erbium bound to the LBT, which is co-expressed with a membrane protein. Red represents zinc in the cell. At NSLS-II, the cell was imaged with a sub-15 nm x-ray beam, which is the highest resolution x-ray fluorescence tomogram of a biological cell ever collected. Image credit: Journal of the American Chemical Society 142 (5), 2145-2149 (2020)
The Science
Lanthanide binding tags (LBTs) were used to image an individual bacterial membrane protein in 3D with a sub 15-nm x-ray beam.
The Impact
The combination of LBTs and x-ray microscopy has the potential to become a widespread tool for imaging individual proteins in whole cells and tissues at the resolution of the cell membrane and subcellular organelles.
Summary
Lanthanide binding tags (LBTs) are short peptides comprised of a nanomolar-a?nity lanthanide-binding domain. LBTs can be easily genetically encoded into a target protein’s sequence and their small size minimizes deleterious e?ects on target protein expression, function, or transport within the cell. The high a?nity of LBTs for lanthanides enables experiments to be performed at low concentrations, thereby mitigating toxic e?ects.
In this study, scientists demonstrated successful 3D x-ray fluorescence tomography of two LBT−protein conjugates, a membrane protein (OmpA) and a soluble protein (ubiquitin). This approach enables the visualization of LBT-tagged proteins while simultaneously measuring the elemental distribution in cells at a spatial resolution necessary for visualizing cell membranes and eukaryotic subcellular organelles.
Using a sub-15 nm beam at the Hard X-ray Nanoprobe (HXN) beamline at the National Synchrotron Light Source II (NSLS-II), the researchers obtained the highest resolution x-ray fluorescence image of a biological cell ever achieved. NSLS-II is a U.S. Department of Energy (DOE) Office of Science User Facility located at DOE’s Brookhaven National Laboratory.
As the conjugation of LBTs and proteins follows standard molecular biology protocols, it is possible that x-ray fluorescence tomography with LBTs could become a widespread tool for 3D imaging of protein distribution and localization within cells with a high spatial resolution while simultaneously obtaining trace element distribution. Moreover, the long penetration depth of x-rays enables nanoscale visualization of the internal structure of cells in 3D, allowing for the extension of the method to tissues.
Download the research summary slide
Related Links
Feature Story: “Cell Membrane Proteins Imaged in 3D”
Contact
Tiffany Victor
NSLS-II, Brookhaven National Laboratory
tvictor@bnl.gov
Publication
T. W. Victor, K. H. O’Toole, L. M. Easthon, M. Ge, R. J. Smith, X. Huang, H. Yan, Y. S. Chu, S. Chen, D. Gursoy, M. Ralle, B.a Imperiali, K. N. Allen, and L. M. Miller. Lanthanide-Binding Tags for 3D X-ray Imaging of Proteins in Cells at Nanoscale Resolution. Journal of the American Chemical Society 142 (5), 2145-2149 (2020). DOI: 10.1021/jacs.9b11571
Funding
This work was supported by the United States Department of Energy, O?ce of Biological and Environmental Research, as part of the “Environment Sensing and Response” Scienti?c Focus Area of the BER Genomic Science Program. T.W.V. was partially supported by the National Institutes of Health T32 Grant 5T32GM092714 and a Director’s Postdoctoral Fellowship at NSLS-II. B.I. and K.N.A. were supported by NSF MCB-1615252 and MCB-1614608, respectively. This research used beamline 3-ID (HXN) at NSLS-II, a U.S. Department of Energy (DOE) O?ce of Science User Facility operated for the DOE O?ce of Science by Brookhaven National Laboratory under Contract No. DE-SC0012704. This research used the BNP beamline at APS, a U.S. DOE O?ce of Science User Facility operated for the DOE O?ce of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
2020-17491 | INT/EXT | Newsroom