Toward Collaborative Scientific Computing
Brookhaven Lab's Scientific Data and Computing Center is developing software tools and enabling technologies to facilitate more effective collaboration among scientists applying computational techniques to solve a variety of scientific problems
September 11, 2019
 enlarge
enlarge
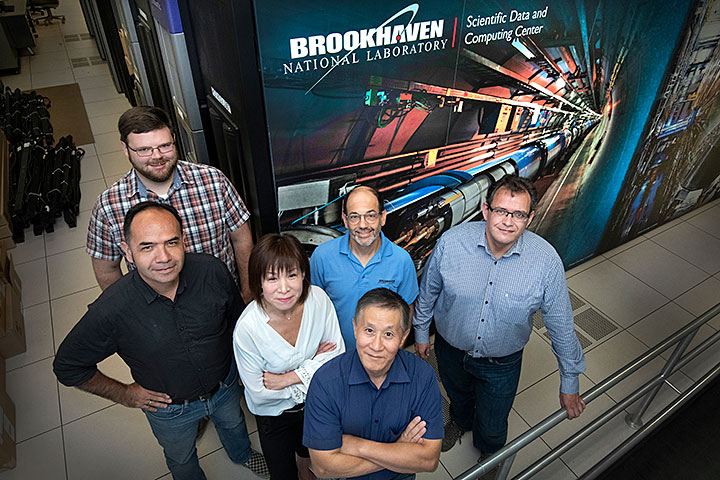
(Front row) Tony Wong, (middle row, from left) Carlos Fernando Gamboa, Mizuki Karasawa, Ofer Rind, Benedikt Hegner, (back row) William Strecker-Kellogg, and (not pictured) John Hover are provisioning a suite of collaborative tools for scientific computing at Brookhaven Lab's Scientific Data and Computing Center.
Billions of people use the World Wide Web to access information and communicate over the Internet. Yet the original idea behind the web, which was invented at the European particle physics laboratory CERN in 1989, was to enable scientists around the world to easily share data from their experiments. Scientific collaboration has only intensified in the modern era of big data, in which experiments are often run at large-scale state-of-the-art facilities through international collaborations that generate larger and more complex data sets at a faster pace than ever before.
In order to derive insights that can lead to scientific breakthroughs, scientists need computing resources to store, manage, analyze, and distribute their data. They also need methods to collaborate—for example, to communicate with each other, organize meetings, search databases, co-develop software code, videoconference, and share documents—while meeting certain requirements such as security.
“The advent of large scientific collaborative efforts in the past decade has ushered in an era in which thousands of geographically dispersed scientists need to work with each other seamlessly,” said Tony Wong, deputy director of the Scientific Data and Computing Center (SDCC), part of the Computational Science Initiative (CSI) at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory.
A center for scientific computing support
Historically, the SDCC has provided a centralized data storage, computing, and networking infrastructure for the nuclear and high-energy physics (HEP) communities and recently has been expanding this infrastructure to newer data-intensive areas, including photon science. For the past year, the SDCC has been extending its support by provisioning a suite of collaborative tools for scientific computing.
“The collaborative tools are not the core software code that scientists employ for their research,” explained Wong. “Rather, they are tools to enable scientists to better work together. They represent a collection of open-source computer software packages that we are reconfiguring to use within the environment at SDCC to support the needs of our expanding user community.”
The SDCC team is currently in the process of adapting several tools:
- Invenio, an open-source software framework for building large-scale repositories to index, store, and curate data, software, publications, presentations, and other content
- Indico, an event management system for scheduling meetings, conferences, and workshops
- BNLBox, a cloud storage service for accessing and sharing large-scale scientific data
- Gitea, a repository hosting software for managing proprietary software code and unpublished scientific papers
- Jupyter, an application for interactive scientific analysis that contains “notebooks” for combining software documentation, code, text, and images
To make such collaboration feasible, the SDCC team is simultaneously developing enabling technologies for user authentication and federated identity management. These vetting processes are required for many of the web-based collaborative tools to safeguard intellectual property and sensitive information.
“Before we deployed these modern collaborative tools at SDCC, we supported a large number of websites with single-sign-on (SSO) technology,” explained SDCC technology architect Mizuki Karasawa, who is developing the authentication and authorization infrastructure. “With SSO, when you try to access a particular website, you are directed to a central identity provider, which prompts you to enter your username and password. Once you are authenticated, you are directed back to the original website, and your identity gets transferred over to all of the services within the website that you are authorized to use—similar to how once you are logged into Gmail, you can access the other Google services without reentering your username and password.”
The current challenge is that many of the collaborative tools that SDCC now offers require newer SSO protocols.
“The traditional SSO enabled at SDCC is no longer applicable,” said Karasawa. “We need to invest in modern SSO technologies.”
Another challenge is that some of the collaborative tools require multifactor authentication (MFA)—a method of confirming users’ claimed identities by using a combination of two different factors: something the users know, something they have, or something they are. One example of MFA requires the user to enter a password and then accept a “push” notification to a personal device such as a cell phone. To address this challenge, Karasawa investigated MFA frameworks using the open-source identity and access management software Keycloak.
In conjunction with this effort, SDCC senior technology architect John Hover is providing enabling technologies for federated user management. Traditionally, users requesting access to SDCC resources and services have had to register with SDCC for an account. But many of the scientists who would make use of the collaborative tools SDCC offers are from universities and other external institutions.
Federated identity management is based on a mutual trust relationship between institutions, or federated partners. When users affiliated with a federated partner request access at another federated partner institution, they enter their username and password from their home institution. Once the home institution verifies the credentials, users are authenticated.
“Brookhaven is part of an identity federation called InCommon, which has about 1,000 members representing research and educational institutions,” said Hover. “There’s also identity management software that we are using with Invenio called CoManage, which was produced by the same entities that established InCommon. The goal is to make collaboration more convenient for scientists. The easier it is for people to use the tools, the more likely they will use them to collaborate.”
 enlarge
enlarge
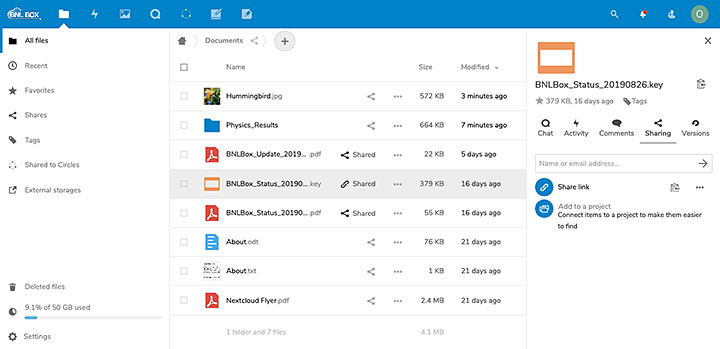
A screenshot of the BNLBox interface, where users can upload and share different kinds of documents.
By providing a data center infrastructure, hosting the collaborative tools, and handling identity authentication in one place, the SDCC offers all of the components necessary for collaborative scientific computing. For example, SDCC’s high storage capacity and BNLBox tool allows scientists to upload and share very large files (which would quickly exceed the free storage space offered by mainstream cloud storage services such as Google Drive or DropBox). These files can be made available to collaborators in a controlled and secure fashion through the identity management frameworks.
According to Ofer Rind, a technology architect at SDCC who has been helping to deploy BNLBox, Indico, and Jupyter, this ability to securely store and share data is one of the advantages of using SDCC services for collaborative scientific computing.
“When you host data in the cloud, you have to be concerned about security,” explained Rind, who is part of SDCC’s RHIC and ATLAS Computing Facility (RACF), which supports the STAR and PHENIX experiments at Brookhaven Lab’s Relativistic Heavy Ion Collider (RHIC) and the ATLAS experiment at CERN’s Large Hadron Collider. “By hosting the data here, we have a higher level of assurance regarding confidentiality, which is an essential working constraint for some scientific programs.”
Use case: nuclear nonproliferation
For example, the National Nuclear Security Administration’s (NNSA) Office of Nuclear Smuggling Detection and Deterrence Science and Engineering Team (SET) generates a variety of programmatic documents, internal reports, and scientific data. The information has traditionally been stored in various locations throughout the national laboratories, such as on local shared drives and on individual hard drives, which are not easily accessible by all team members. To facilitate knowledge management and sharing, SET tasked Brookhaven’s Nonproliferation and National Security Department (NNSD), with support from CSI, to develop an advanced data repository.
After conducting an evaluation of existing institutional repository software, the team selected the Invenio software framework to build the SET repository.
“Invenio, developed by CERN, is an open-source framework for the management of large-scale digital assets,” explained Uma Ganapathy, the lead software engineer on the project and an advanced applications engineer in CSI’s Center for Data-Driven Discovery (C3D). “The SET repository uses the underlying infrastructure support provided by Invenio for data storage, retrieval, and search. For the nonproliferation program, the repository has been made private with restricted access. Several other features have been added to the application for customized data models for storage and more refined access control and search. We plan to extend the tool to perform document reviews, approvals, and data analysis.”
After intense development and testing, the team released the first version of the SET repository—which is hosted within the computing infrastructure at SDCC—in early July. After a two-week trial run, the repository was officially launched.
“We are pleased that we can deliver to NNSA a repository with capabilities more advanced than any others currently being used by NNSA,” said principal investigator Warren Stern, a senior advisor in NNSD. “This repository will support NNSA’s aggressive effort to detect nuclear smuggling by using tested modern data tools, and bring capabilities developed in the scientific community to the nuclear security community.”
Use case: materials synthesis
Ganapathy is also developing an Invenio-based repository for the Next-Generation Synthesis Center (GENESIS), a new DOE Energy Frontier Research Center directed by Stony Brook University professor John Parise that was created to accelerate the discovery of new materials. The DOE scientists, university faculty, and students and postdocs involved in this project will use modern characterization tools such as photon sources—including Brookhaven’s National Synchrotron Light Source II (NSLS-II)—along with data science tools, to study, predict, and eventually control the chemical reaction pathways that govern material synthesis.
According to Line Pouchard, the Brookhaven principal investigator on the project, SDCC provides scalability for GENESIS—that is, enough space to store not only the collected material sample data but also metadata about the parameters under which experiments were carried out and the methods used to perform data reconstruction.
“Such data history or traceability—what we call provenance—is important for reproducing, validating, and comparing results,” said Pouchard. “Another advantage of using SDCC’s services is their expertise in storing, safeguarding, and distributing data. The team at SDCC worked with Brookhaven’s Information Technology Division to implement federated identity management so that people who are vetted in the project can access the repository by using their university credentials. Otherwise, everyone, including grad students who are always coming and going, would have to request guest appointments.”
Users from other InCommon institutions can log in and use the GENESIS Invenio application without Brookhaven computing accounts. And CoManage allows the GENESIS project leaders to manage their own membership without SDCC staff involvement.
 enlarge
enlarge
Brookhaven is provisioning a suite of software tools and enabling technologies to enable collaboration among geographically dispersed scientists.
Currently, Ganapathy is improving Invenio based on detailed user feedback. For example, users requested a particular archiving hierarchy, or file arrangement. In addition, they asked for the ability to perform searches that are specific to materials science, such as sample composition.
In the future, the Invenio framework will be extended to different scientific communities.
“Ideally, what we want to have is an Invenio-based research data management platform that is homogenous and flexible enough for different types of use cases,” said SDCC senior technology engineer Carlos Fernando Gamboa. “Working toward this goal, we first deployed Zenodo/Invenio (a research data management platform developed at CERN) as a test pilot in two communities (nuclear nonproliferation and materials science). By interacting with the Invenio framework and testing its capabilities, these communities built their own digital repositories to meet their specific needs (SET and GENESIS). The SDCC is now working with CERN and ten other multidisciplinary and commercial institutions to build a research data management platform called InvenioRDM. One community already looking to us for such a content management solution is nuclear and particle physics.”
Use cases: big physics experiments
Besides content management, other collaborative tools for scientific computing are in demand by large nuclear and particle physics experiments. Brookhaven Lab is involved in three such experiments: the STAR and sPHENIX nuclear physics experiments at RHIC, and the HEP experiment Belle II at the SuperKEKB accelerator in Japan. Each experiment is trying to seek out new physics within subatomic particles that could help us understand mysteries related to the early universe.
For instance, scientists in the STAR collaboration are studying the quark-gluon plasma—a very hot, dense “soup” of matter’s fundamental building blocks that existed only a few millionths of a second after the Big Bang. The STAR detector, located at RHIC, runs for about six months of the year for 24 hours a day. During STAR runs, scientists work in shifts to monitor RHIC collisions. The recorded collision events must be logged and passed along to multiple shift crews. To perform these tasks, scientists need to attach and save many documents to the shift log. Providing a seamless interface that allows users to manually attach event pictures taken with a smartphone, screenshots of a monitoring system, or an updated user manual is not always easy, especially considering that some of the experts involved are located offsite at other institutions.
“We thought we could do what people do on social media: snap a picture and upload it,” said STAR physicist Jerome Lauret, who has participated in gathering the requirements for collaborative tools. “While we could develop such an interface, the simpler way is to use BNLBox to add pictures from anywhere and anytime and then add them to our shift log. We can also use BNLBox to exchange documents. The advantage over other box commercial tools is that we own the storage and hence the content. This feature alleviates the security concerns some may have in using such distributed storage in our environment.”
Lauret continues to work closely with the SDCC team to ensure that BNLBox comes to a full production-ready quality.
In addition to requiring data storage and access, scientists need ways to manage meetings. For example, the nearly 250 scientists that make up the sPHENIX collaboration—which will expand on discoveries made by STAR and the predecessor PHENIX experiment to study how the QGP’s properties arise from the interactions of quarks and gluons—regularly meet at workshops, conferences, and other gatherings to discuss their progress and the path forward. From booking rooms and allotting speaker time slots, to uploading and archiving meeting materials such as agendas and presentations, running these events involves a lot of coordination from a logistics standpoint.
Indico, the event management system that was developed at CERN, has long been the primary tool for this task at Brookhaven. However, the latest versions of this software lack some search and videoconferencing capabilities that are highly sought after among the Brookhaven user base. Seeing this pressing need, the SDCC team formed a collaboration with developers at CERN and DOE’s Fermilab to develop and implement a set of plugins that will provide these functionalities.
Scientists involved in the sPHENIX collaboration are also using Gitea to manage proprietary codes and unpublished papers. Unlike the versions of this tool that offer public repositories, such as Github and Gitlab, Gitea enables them to create private repositories with restricted access.
For both Indico and Gitea, the SDCC team is implementing an SSO functionality.
“SDCC used to mainly be a computing center that provided the computing resources to handle a very large amount of data,” said Chris Pinkenburg, a nuclear physicist in the sPHENIX group. “The new collaborative tools that SDCC is now providing are highly appreciated because they help us efficiently run such a big data experiment.”
Belle II is another big data experiment that is benefitting from the services offered by SDCC. Scientists involved in this HEP collaboration are trying to understand the difference between matter and antimatter by colliding electrons and positrons and looking for extremely rare processes that occur during the collisions.
“In our current understanding of the universe starting with the Big Bang, equal amounts of matter and antimatter should have been created,” said Brookhaven Lab physicist Paul Laycock. “When combined, these particles annihilate each other. So how come there is a difference between matter and antimatter such that there remains some matter today out of which the whole universe was made, instead of an empty, black nothingness? How come we’re here at all?”
Answering these questions requires specialized software to analyze and fit the physics data generated from the collisions. The SDCC team is configuring Jupyter so that it can access the Belle II software.
“In the past, people used to open their favorite software editor and make changes in the code itself and then they would run that code,” said Laycock. “In contrast, Jupyter allows us to combine Belle II software documentation and execution in a single platform. This combination makes Jupyter a great educational tool for showing scientists how to use the Belle II software and for collaborating on software development.”
The SDCC team set up this interactive computing environment in such a way that users not only have access to the service itself but also to computing resources at SDCC.
“Jupyter was originally deployed as a tool for ATLAS but has since found a variety of other interested users at the Lab,” said SDCC technology engineer William Strecker-Kellogg. “We combined existing and custom code to integrate the Jupyter service with resources at our data center. When users have a Jupyter session open in their web browser, they remotely have access to all of their data and can run analyses on SDCC’s high-performance and high-throughput computers.”
Use cases: nanomaterial characterization
Strecker-Kellogg is also providing Jupyter to users of two kinds of software developed at the Center for Functional Nanomaterials (CFN): one to analyze x-ray scattering nanostructure data generated at the Soft Matter Interfaces beamline at NSLS-II (CFN is a partner user on this beamline), and another to detect nanoparticles in environmental transmission electron microscopy (TEM) images.
“For a number of instruments here at the CFN and at NSLS-II, the size and complexity of the datasets are growing dramatically,” said Mark Hybertsen, leader of the CFN Theory and Computation Group. “Thus, we are making significant investments in developing automated methods to extract information out of these datasets.”
Consider the environmental TEM at the CFN. This instrument has a high-speed camera, which can record a video of a sample as its evolving over time under some conditions as fast as one frame every millisecond. So, a video with more than 100,000 frames could quickly be generated. Say there are a few hundred nanoparticles in each frame. Maybe some of these particles are getting bigger or smaller, or they are moving around.
“It would be impossible to measure the particle sizes and track them in the images by hand,” said Hybertsen. “That’s why a CFN team is developing particle recognition and tracking software.”
However, the datasets resulting from such TEM experiments are extremely large—in the terabyte range (about 50,000 trees would be needed to produce enough paper to hold the equivalent of one terabyte of data)—and thus require powerful computing resources for storage and analysis.
The SDCC team is building a framework for scientists who use Brookhaven facilities to get access to their data and software tools—such as those designed for TEM data analysis—remotely from their home institutions. This framework supports Jupyter tools, which in turn support graphical user interfaces and other features that make it much easier for users unaccustomed to doing scientific computing themselves.
 enlarge
enlarge
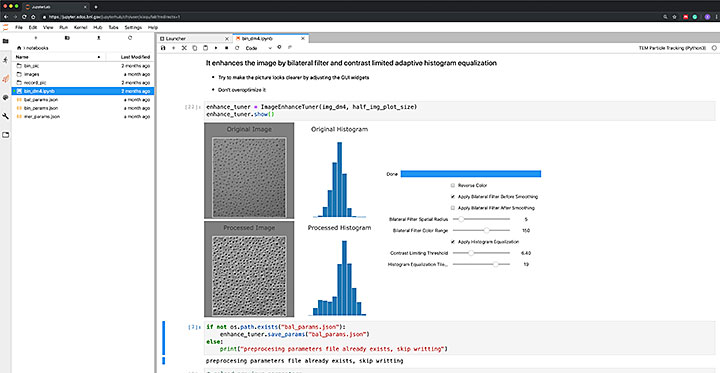
A screenshot of a Jupyter notebook to set up the parameters for segmenting images of nanoparticles recorded by a transmission electron microscope. The widgets include those that allow users to adjust the preprocessing parameters to improve image quality and see the results interactively. The digitalized data will be used to analyze the process of particles evolving and diminishing.
Scientists at NSLS-II are also using Jupyter. Strecker-Kellogg is collaborating with NSLS-II computational scientist Daniel Allan on a Jupyter extension that will allow users to sync and share live notebooks from within one session. In the future, the hope is that NSLS-II data—which is currently stored in the facility and involves local storage at each beamline—will be transferred over to SDCC. Such centralized data management would allow for a more streamlined use of Jupyter.
“NSLS-II was one of the earliest institutions to try JupyterHub, the multi-user server for Jupyter notebooks,” said Allan. “We have been maintaining our own deployment customized to our users’ needs for several years. Responding to our users’ needs, we have developed tools for browsing available computation environments and defining new ones, and sharing notebooks and environments—both from staff to users and between users. We have shared these tools with Jupyter’s open-source community, and other institutions have contributed their improvements.”
Now, NSLS-II is in the process of migrating from the NSLS-II JupyterHub to the SDCC-managed lab-wide JupyterHub deployment.
“We look forward to expanding and developing our customizations to benefit the whole lab,” said Allan. “In turn, NSLS-II users will benefit from the expertise, stability, and infrastructure that SDCC provides. Users analyzing data from their home institutions will have access to dedicated data analysis resources located at SDCC, rather than sharing individual beamline servers with users who are on site acquiring data. This approach keeps users who are analyzing data and those who are acquiring new data out of one another’s way and enables us to consider separately what resources they require.”
Future of collaboration support
Going forward, the SDCC will continue to increase the quantity of collaborative tools that enable scientists to share information securely and collaborate productively. They will also extend the availability of these tools to other scientific user communities. An emphasis will be placed on tools that expedite the process from data gathering through data analysis and scientific discovery, to data and document preservation. For example, one service that SDCC is currently looking into is group chat software.
Staff from SDCC will present their latest activities on collaborative tools at several upcoming conferences and workshops, including Federated Identity Management for Research (September 12), HEPiX (October 14–18), and the International Conference on Computing in High Energy and Nuclear Physics (CHEP) (November 4–8).
“Collaborative tools are the glue that enables scientists scattered around the globe to work effectively,” said Wong. “As an example of their importance and relevance, collaborative tools are listed explicitly as a topic in prominent scientific gatherings such as CHEP. The SDCC is committed to providing the services and enabling technologies that make collaboration possible.”
NSLS-II, RHIC, and CFN are all DOE Office of Science User Facilities.
Brookhaven National Laboratory is supported by the U.S. Department of Energy’s Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit https://energy.gov/science.
Follow @BrookhavenLab on Twitter or find us on Facebook.
2019-16638 | INT/EXT | Newsroom