Following the Data Trail to Accelerated Discovery
A team of computational scientists, software engineers, and physicists is developing software that will keep track of sample descriptions, experimental conditions, and data analysis methods so scientists can interpret, validate, compare, and reproduce results—and eventually automate their research
October 4, 2019
 enlarge
enlarge
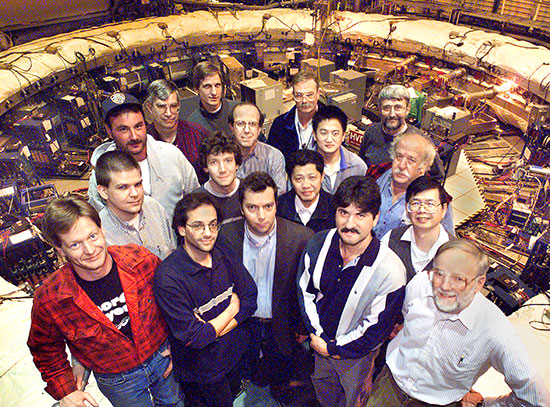
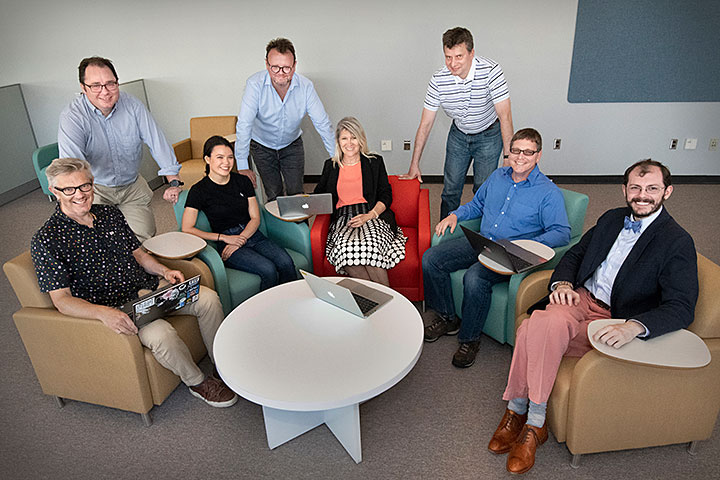
(Left to right) Simon Billinge, Stuart Campbell, Aiko Hassett, Eli Stavitski, Line Pouchard, Pavol Juhas, Hubertus van Dam, and Christopher Wright (not pictured: Gilchan Park) are developing software to record the trail of data capture and transformation. This Dynamic Provenance System is being piloted at two beamlines at Brookhaven Lab's National Synchrotron Light Source II.
Knowing where you come from is a natural curiosity—one that has led to the entirely new industry of ancestry testing. With a saliva sample, companies such as Ancestry.com and 23andMe can automatically compare your genetic information to that of other people from all around the world who have also submitted a DNA kit. Millions of consumers have already added their DNA to these companies’ databases, some of which also contain extensive historical documents, family trees, and images for genealogical research. This vast trove of information only keeps growing as more people join the online communities and new records are added.
This desire to trace origins also extends to data itself. In computer science, the ability to record the origin and history of data is known as provenance.
“Provenance is the record of data lineage and software processes operating on these data that enable the interpretation, validation, and reproduction of results,” explained Line Pouchard, a senior researcher at the Center for Data-Driven Discovery (C3D), part of the Computational Science Initiative (CSI) at the U.S. Department of Energy’s (DOE) Brookhaven National Laboratory.
Currently, Pouchard is leading an effort focused on enhancing scientific experimentation through provenance via a framework called the Dynamic Provenance System (DPS).
For scientific experiments, provenance encompasses descriptions of samples, experimental procedures and conditions, and data analysis methods. The ability to record these metadata and scientific workflows is especially critical in today’s era of big data, where researchers are faced with large, diverse data from complex, dynamic, and streaming heterogeneous sources. In order for scientists to derive insights that lead to discoveries, they need to know how data were generated and transformed from their original state to produce the final results.
An existing data infrastructure
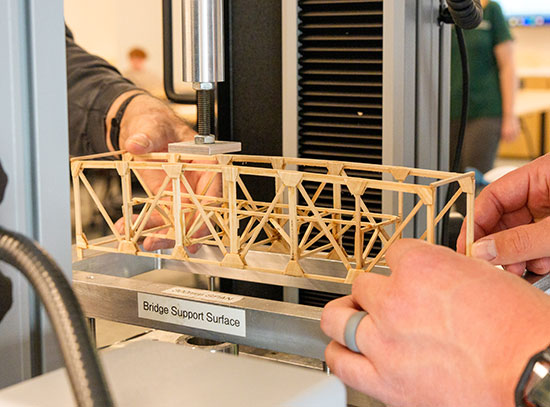
Consider the National Synchrotron Light Source II (NSLS-II)—a DOE Office of Science User Facility at Brookhaven Lab where scientists use ultrabright x-ray light to reveal the atomic and electronic structure, chemical composition, and magnetic properties of materials. By 2020, NSLS-II is expected to produce more than 20 petabytes of experimental data. For reference, 1.5 million CD-ROM discs would be required to store a single petabyte. Currently, 28 experimental stations, or beamlines, are in operation at NSLS-II. That means 28 separate experiments can be ongoing at once. Once fully built out, NSLS-II could accommodate up to 60 beamlines.
Traditionally, each beamline has provided its own data acquisition and analysis tools. But recently, Brookhaven’s Data Acquisition, Management, and Analysis (DAMA) Group developed a collection of software tools named Bluesky to streamline the process of data collection and analysis across different beamlines and even other light sources. This open-source software has also been deployed at three other DOE Office of Science User Facilities: the Advanced Photon Source at Argonne National Lab, the Advanced Light Source at Berkeley National Lab, and the Linac Coherent Light Source at SLAC National Accelerator Lab.
“Bluesky allows us to store the entire data trail, from the birth of the sample and its characteristics—for example, who made it, how and when it was made, its chemical composition, and its physical state—to experimental conditions and instrument calibrations, to the software parameters used to analyze the data, to the tangible scientific results generated by analyzing the raw data,” explained Eli Stavitski, lead scientist at NSLS-II’s Inner Shell Spectroscopy (ISS) beamline. “In order to support reproducible science, all of these metadata have to be tracked by a provenance system.”
Provenance for photon science
To help boost the data and computing infrastructure at NSLS-II, Pouchard’s team has been developing provenance software that leverages Bluesky.
“Our DPS enables searching across the beamlines, provides statistics on samples studied at NSLS-II, and brings external data sources right to scientists’ fingertips during their experiments,” Pouchard explained. “These sources include crystallography databases and information extracted from the scientific literature.”
In addition to Pouchard and Stavitski, the team members are C3D computational scientist Pavol Juhas and postdoctoral research associate Gilchan Park; application architect Hubertus van Dam of CSI’s Computational Science Laboratory; DAMA group leader Stuart Campbell; Simon Billinge, a physicist in Brookhaven’s Condensed Matter Physics and Materials Science Department and a professor of materials science and engineering and applied physics and applied mathematics at Columbia; CJ Wright, a recent PhD graduate of Columbia and lead software engineer for the Billinge Group; and Aiko Hassett, a computer science major at Middlebury College who participated in DOE’s Science Undergraduate Laboratory Internships program in summer 2019.
The team has deployed a prototype version of this provenance software at two NSLS-II beamlines: ISS and X-ray Powder Diffraction (XPD).
“ISS and XPD are what we call high-throughput beamlines,” said Campbell. “These beamlines are capable of measuring up to hundreds of samples in a single day. While Bluesky provides a framework for storing the metadata from these experiments in databases, we need provenance “dictionaries,” or a set of metadata schemas, that are customized for specific scientific techniques and samples.”
 enlarge
enlarge
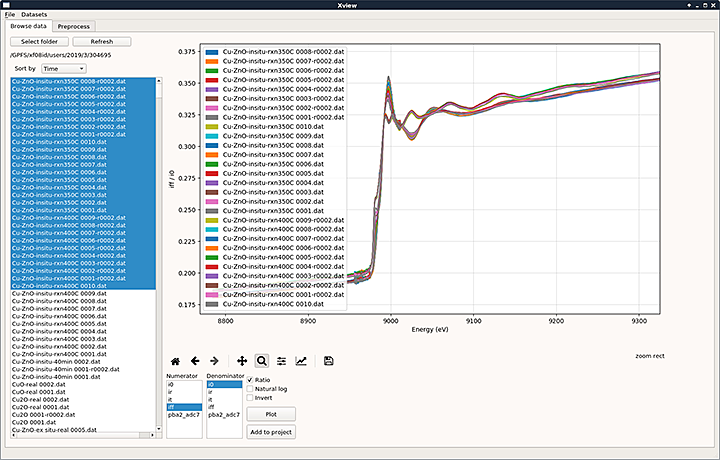
A screenshot of the data acquisition and visualization software—which uses the code developed by Pouchard's team—at the ISS beamline. The curves are a selected subset of x-ray absorption spectra for a copper zinc oxide (Cu-ZnO) sample recorded at ISS, with the different colors discriminating between curves. X-ray absorption spectroscopy is a method to characterize the local atomic and electronic structure of materials.
Metadata schemas provide an overall structure for organizing and formatting the metadata in a standardized way. Without well-defined schemas, retrieving relevant information from the database would be difficult.
“The DPS software allows users to search for metadata on samples and previously performed experiments and to replay analysis workflows,” said Pouchard.
For previous generations of light sources, scientists traditionally analyzed their scientific results back at their home institutions. But finding out after the experiment that there was a problem with the experimental setup or the data were insufficient to make any conclusions is not very helpful, especially considering that beam time is limited.
“We would like to enable data analysis while the scientists are doing their experiments at the beamlines so they can make adjustments if necessary,” said van Dam. Such streaming analysis requires computational power and networks, as well as optimized analysis codes.”
“Parallel computing—farming out computations in many nodes—can speed up very-high-intensity calculations,” explained Wright. “In the context of streaming data, executing parallel computing is challenging because the computations from subsequent data points can complete before preceding ones. If the data are not processed in the right order, then the accuracy of the results can be significantly impacted.”
This on-the-fly analysis could enable scientists to compare their results with preexisting data sets to find potential reference structures. Similarly, while the scientific literature contains a wealth of information, it is not very useful if it cannot be parsed in an efficient way that provides relevant answers to very specific questions. According to Pouchard, this problem is why DPS also aims to provide scientists with timely, relevant information extracted from the scientific literature. To help with this task, Park has been developing a text and data mining system based on natural language processing—a branch of artificial intelligence that involves designing computer algorithms that can understand human language.
“Within the past 50 years, thousands and maybe millions of experimental results similar to what we’re doing here at NSLS-II were reported in scientific journals,” said Stavitski. “With the ever-growing number of journals, tracking of all of these papers has become impossible for a human to do. The text and data mining system will not only allow us to more quickly find previously published results for comparison but also to extract pieces of provenance schemas in a form that can be fed into machine learning (ML) algorithms. What comes out from the beamline is a set of numbers, from which figures are created to include in scientific publications. In order for ML to take advantage of these data, the figures need to be transformed back into numbers.”
Toward autonomous experiments
If the DPS prototype targeting a small subset of beamlines proves to be successful, the idea is to extend it throughout the rest of NSLS-II, as well as other facilities where NSLS-II infrastructure has been deployed. According to Stavitski, one of the challenges in a facility-wide implementation within NSLS-II is making schemas that are universally applicable across all beamlines.
“What we need is for the schemas to work like Lego pieces that for a particular type of experiment on a particular beamline, you select the ones that make sense for that technique and sample type,” said Stavitski. “As you move to a different sample or technique, you select a different set of schemas.”
As Campbell notes, this flexibility would be particularly helpful for automated multimodal analysis.
“When multiple measurements with different beamlines are taken on the same sample, future automated analysis would be able to parse these data to provide a more complete picture of the science of that sample,” explained Campbell.
“With our ability to capture data using so many different (but complementary) techniques, we have to delegate some of the analysis to intelligent computer systems,” said Stavitski. “It is no longer feasible for a human to keep up. But machine-based analysis is only possible if every piece of information has been tracked.”
"Because most of ML is done on labeled, structured data, you need to have databases filled with high-quality metadata,” added Billinge. “So, for example, if you have a bunch of x-ray diffraction images stored in a database, you can do ML by training the algorithm to recognize images that are coming from a liquid versus crystal sample. This is where a computational infrastructure becomes important because it enables the capture of the metadata needed for ML. We haven’t had that in place until now.”
Now that Pouchard’s team has demonstrated the generalizability of the DPS on two different NSLS-II beamlines, they would now like to show that it is scalable. Pouchard noted that such scalable provenance systems are needed across many domains, including in beamline experiments, computational experiments, and performance analyses of computer systems. Part of this scalability will involve developing provenance-tracking capabilities for combined experimental and simulation data. Simulations can complement experiments by predicting what to expect and revealing the atomic-scale processes underlying observations—for example, the changes in the chemistry of electrode materials that occur as a battery charges and discharges.
“Another layer of complexity is involved because there are completely different data sets (measured versus calculated), each with varying degrees of accuracy,” said van Dam. “We need to figure out how to compare them in a valid way. One of the issues is computational reproducibility in ML. Computational reproducibility covers a whole range of thresholds, including accuracy and uncertainties. When you do ML with scientific data, the methods are not deterministic; the outputs of the computations are based on the parameters you have chosen.”
The team is also interested in exploring whether a similar provenance system can be used at other facilities with extremely high data acquisition rates, such as the Center for Functional Nanomaterials—another DOE Office of Science User Facility at Brookhaven.
“Ultimately, provenance will enable better science, as it will increase confidence in the methods used to obtain a particular result,” said Pouchard. “It will also enable researchers to reproduce results and help them learn from past experiments.”
The research is funded by Brookhaven’s Laboratory Directed Research and Development program, which promotes highly innovative and exploratory projects.
Brookhaven National Laboratory is supported by the U.S. Department of Energy’s Office of Science. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit https://energy.gov/science.
Follow @BrookhavenLab on Twitter or find us on Facebook.
2019-16668 | INT/EXT | Newsroom









