Resolving the Internal Structure of Nanoparticle Dimers Linked by DNA
January 15, 2013
What Is The Scientific Achievement?
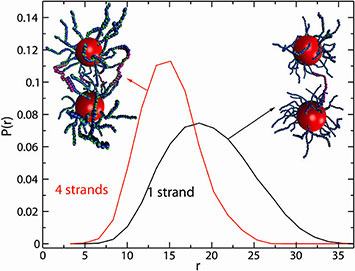
Figure A: Calculated distribution of interparticle separation, r, for dimers linked by either one or four DNA chains. The average separation decreased with increasing number of particle-connecting chains.
Light and x-ray scattering experiments have revealed the structure of nanoparticle dimers linked by flexible DNA chains. These dimers were basic units in a multi-scale,hierarchical assembly, and served as a model system to understand DNA-mediated interactions, particularly in the non-trivial regime when the nanoparticle and DNA links were comparable in size. We found that the interparticle separation within the dimer was primarily controlled by the number of linking DNA. We summarized our findings in a simple model that captured the interplay of the number of DNA bridges, their length,the curvature of the nanoparticle, and the excluded volume effects. We demonstrated excellent agreement of our analytical model with both our experimental and computational results, without use of free parameters in the model.
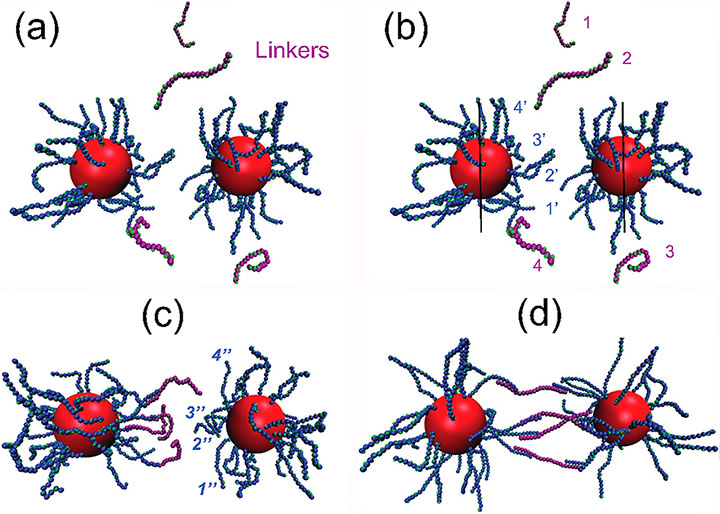
Figure B: Simulation protocol for dimer formation of two DNA-coated NPs connected by four linkers. (a) Two DNA-coated NPs and four linkers (pink strands) are located at random positions. (b) Four strands of one hemisphere of each NP are randomly selected. (c) Hybridization occurs between the linkers and strands of the left NP. (d) Hybridization with second NP followed by equilibration.
Why Does This Matter?
As building blocks in multi-scale assemblies, DNA-linked nanoparticle dimers are excellent model systems to understand chain-mediated interactions between particles. The results of this study can serve to guide the design of precisely controlled distances within nanoscale clusters; such control is imperative for energy transfer and bio-detection functionalities, among other applications.
What Are The Details?
- CFN Capabilities: X-rays cattering at CFN/NSLS X9 Endstation, UV-Vis, Dynamic Light Scattering, and TEM allowed for investigations of the self-assembly processes of the DNA linked dimers. CFN/NSLS X9 Endstation and Dynamic Light Scattering also facilitated in-situ probing of the interparticle distance within the dimers.
- When nanoparticles and DNA chains are of comparable size, the inter-particle separation within a dimer deviate from a free-chain behavior: in that case, the well-established dependence on the chain length is suppressed due to the presence of multiple DNA connections between the highly curved surfaces of the nanoparticles. Our analytical model exhibits excellent agreement with both experimental and computational results.
Publication Reference
Internal Structureof Nanoparticle Dimers Linked by DNA
Cheng Chi,† Fernando Vargas-Lara,‡ Alexei V. Tkachenko,† Francis W. Starr,‡ and Oleg Gang†
† Center for Functional Nanomaterials,Brookhaven National Laboratory, Upton, New York 11973, United States
‡ Physics Department, Wesleyan University,Middletown, Connecticut 06459, United States
ACS Nano 6 (8), 6793-6802 (2012)
http://pubs.acs.org/doi/abs/10.1021%2Fnn301528h
Acknowledgment of Support
Research was carried out at the Center for Functional Nanomaterials, Brookhaven National Laboratory, which is supported by the U.S. Department of Energy, Office of Basic Energy Sciences, under Contract No. DE-AC02-98CH10886. F.V.-L. and F.W.S. thank the NSF for support from grant number CNS-0959856.
2013-3616 | INT/EXT | Newsroom