New Polymer Model Helps Explain the Origins of Life
February 19, 2019
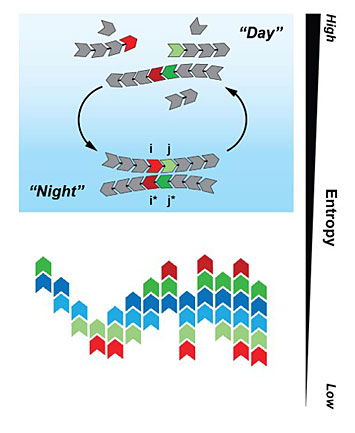
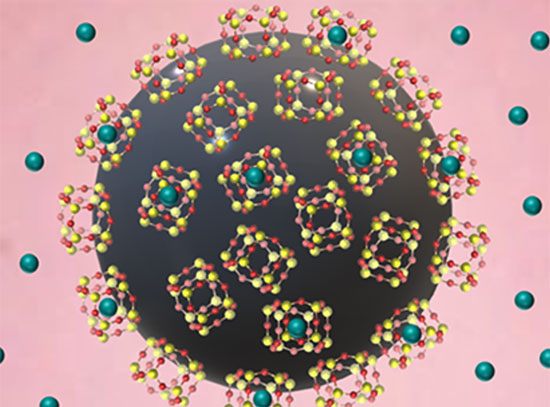
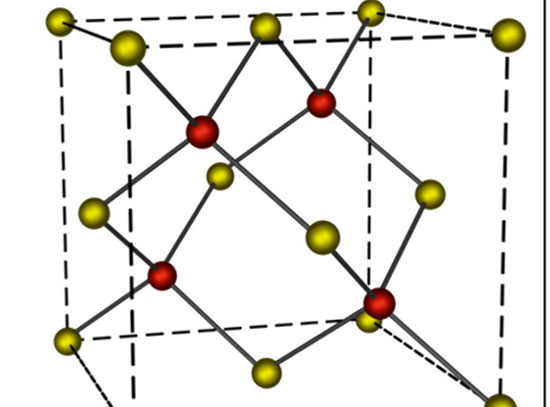
Illustration of the Tkachenko-Maslov model shows different monomers cycling between "day" and "night"
What is the scientific achievement?
CFN staff in collaboration with the University of Illinois at Urbana-Champaign developed a polymerization model that evolves from a "soup" of different monomers into a complex system made from a limited number of polymer segments, which reduces the entropy in the system. Numerical results, supported by mathematical analysis, confirm the survival and extinction process that resembles natural selection, and drives the dramatic decrease of informational entropy.
Why does this achievement matter?
The emergence of life from non-living matter is one of the greatest mysteries of fundamental science. The model could also be applied to make biopolymers and nanostructures.
What are the details?
Understanding the onset of the reduction of information alongside ever-increasing complexity is essential for solving the problem of the origin of life. To study heteropolymers capable of template-assisted ligation based on Watson-Crick-like hybridization, the system is driven off-equilibrium by cyclic changes in the environment, and the population dynamics of sequential pairs of specific monomers within the heteropolymer population (called “2-mers”) was then modelled. While the possible number of such pairs is Z2 (where Z is the number of monomer types), most of them eventually become extinct, leaving no more than 2Z survivors. This leads to a dramatic reduction of the information entropy in the sequence space. The numerical results are supported by a general mathematical analysis of the competition of growing polymers for constituent monomers. This natural-selection-like process ultimately results in a limited subset of polymer sequences. An inhomogeneity in initial conditions allows for a massively parallel search of the sequence space for biologically functional polymers, such as ribozymes. There are also potential experimental implementations of this model in the contexts of either biopolymers or artificial nano-structures.
CFN Capabilities:
The CFN Theory and Computation Facility was used for this collaboration.
Publication Reference
A. V. Tkachenko and S. Maslov, Onset of natural selection in populations of autocatalytic heteropolymers, Journal of Chemical Physics 149, 134901 (2018).
Acknowledgement of Support
The research was partially carried out, and used computational resources, at the Center for Functional Nanomaterials, user facility at Brookhaven National Laboratory under Contract DE-SC0012704 with the U.S. Department of Energy.
2019-14392 | INT/EXT | Newsroom