Bond, DNA Bond. Assembling Ordered Arrays at the Nanoscale
September 30, 2021
 enlarge
enlarge
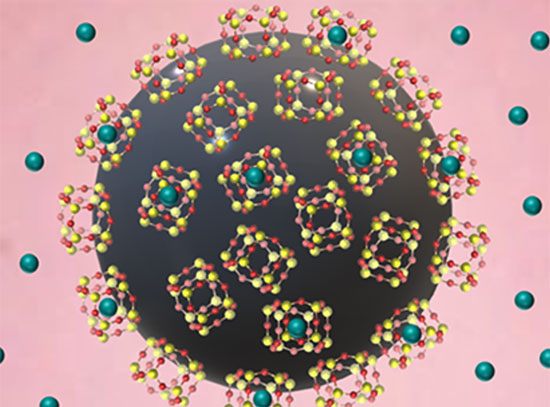
Box-shaped DNA "nanochambers"—which can carry cargo such as nanoparticles—are programmed to assemble into 1D, 2D, or 3D ordered arrays, depending on the prescribed binding mode. Illustration featured on the cover of the Journal of the American Chemical Society.
What is the scientific achievement?
CFN staff led a team of users and collaborators in developing a strategy for folding DNA molecules into hollow cubes with fully prescribed bonds encoded along different directions. By designing specific binding modes, the team programmed the DNA “nanochambers”—which can carry nanocargo—to self-assemble into 1D, 2D, and 3D ordered arrays.
Why does this achievement matter?
Control over the binding properties of nanoscale building blocks is needed to guide their assembly into complex, desired structures for applications including drug delivery, photonics, and catalysis.
What are the details?
DNA origami—inspired by the Japanese art of folding paper in particular ways to make various shapes—has emerged as a powerful technique for the programmable self-assembly of materials. Here, scientists used a long DNA molecule as a scaffold and folded it into a cube. Analogous to packing simple paper crafts into complex shapes, the DNA cubes serve as basic building blocks that can be further assembled into desired nanostructures. However, engineering the assembly of nanoscale objects into complex and prescribed structures requires control over their binding properties. Although much progress has been achieved in scientists’ ability to design complex nano-objects, creating such nano-objects with fully controlled binding modes and understanding their fundamental properties remain a challenge. In this study, CFN users and staff and collaborators from Columbia University and Georgia Tech/Emory University developed a strategy for creating DNA “nanochambers,” hollow cuboid nano-objects whose DNA bonds can be fully prescribed and complexly encoded along their three orthogonal axes. By tuning the directionality of DNA bonds, they built one-, two-, and three- dimensional ordered arrays of DNA nanochambers. The team demonstrated how these nanochambers can host nanoscale cargo such as gold nanoparticles, allowing for the construction of complex organizations of nanocargoes with controlled architectures. Transmission electron microscopy at the CFN Electron Microscopy Facility enabled direct visualization of the DNA nanochambers and their organized arrays, while in situ small-angle x-ray scattering at two NSLS-II beamlines that the CFN is a partner user on through its Advanced UV and X-ray Probes Facility—the Complex Materials Scattering (CMS) and Soft Matter Interfaces (SMI) beamlines—enabled the in-situ characterization of these arrays. The scientists combined these experimental investigations with computational simulation to understand the mechanism of structural formation for different types of ordered arrays and to correlate bond design with assembly processes.
CFN Capabilities
The CFN Electron Microscopy Facility was used to image the DNA nanochambers and assembled arrays. X-ray scattering at the CMS and SMI beamlines, operated in partnership between CFN and NSLS-II, was used to characterize programmed arrays in situ.
Publication Reference
Z. Lin, H. Emamy, B. Minevich, Y. Xiong, S. Xiang, S. Kumar, Y. Ke, and O. Gang, “Engineering Organization of DNA Nano-Chambers through Dimensionally Controlled and Multi-Sequence Encoded Differentiated Bonds,” Journal of the American Chemical Society 142, 41 (2020).
DOI: https://doi.org/10.1021/jacs.0c07263
Acknowledgement of Support
This research was supported by the U.S. Department of Energy, Office of Basic Energy Sciences, Division of Materials Sciences and Engineering under grant DE-SC0008772. X-ray scattering experiments were carried out at the Complex Materials Scattering (CMS) and the Soft Matter Interfaces (SMI) beamlines at the National Synchrotron Light Source II, which are U.S. DOE Office of Science Facilities at Brookhaven National Laboratory under contract no. DE-SC0012704. This research used resources of the Center for Functional Nanomaterials, supported by U.S. DOE Office of Science Facilities at Brookhaven National Laboratory under Contract No. DE-SC0012704. We would like to acknowledge the Imaging Facility of CUNY Advanced Science Research Center for the instrument use and technical assistance.
2021-19234 | INT/EXT | Newsroom