Arbitrary Design of DNA-Programmable 3D Crystals through Symmetry Mapping
January 28, 2026
 enlarge
enlarge
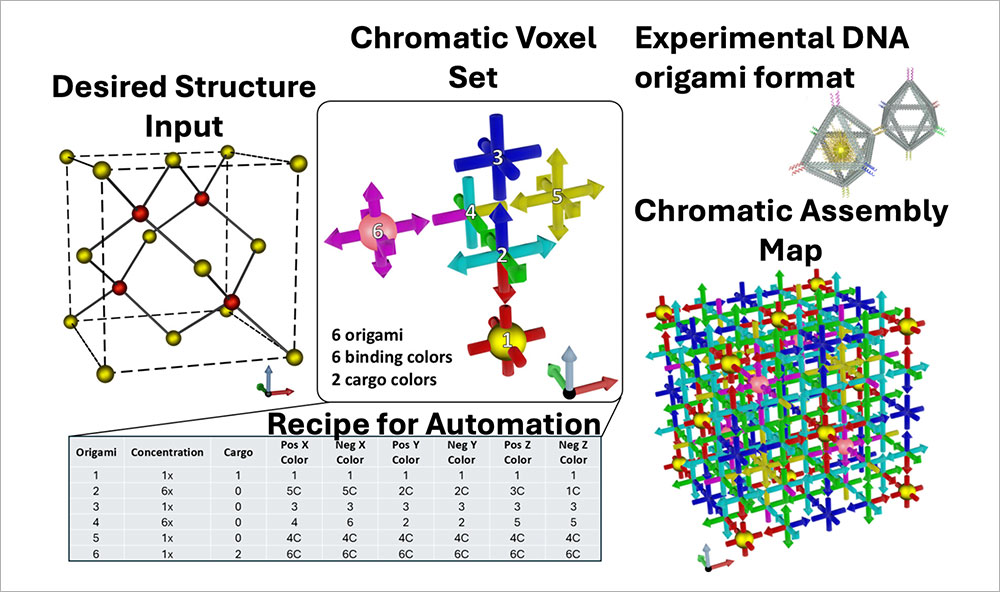
The MOSES mapping algorithm takes a target 3D lattice as input and, via inverse design, outputs a minimal materials design and a color-coded (chromatic) bond map — a complete assembly recipe.
Scientific Achievement
CFN researchers and collaborators developed a symmetry-mapping algorithm, MOSES, that streamlines the design of DNA voxels and can be generalized to patchy particles. These voxels are capable of self-assembling into complex three-dimensional superlattices.
Significance and Impact
This approach reduces the complexity of molecular programming, enabling scalable, modular, and programmable routes to 3D nanostructures and automated materials design.
Research Details
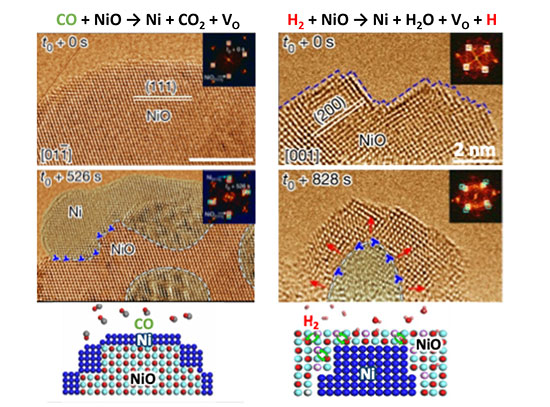
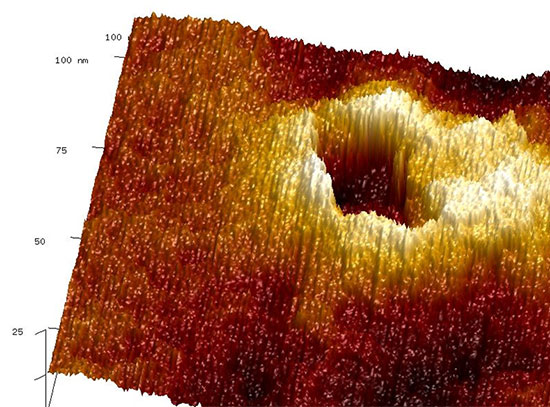
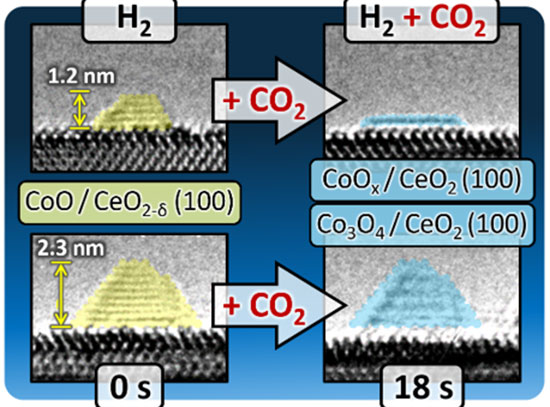
Nanoscale self-assembly offers a powerful route to fabricating intricate architectures beyond the limits of traditional top-down nanofabrication. In particular, DNA nanotechnology has emerged as a versatile and increasingly widely used platform for nanoscale construction because DNA strands can be designed to recognize and bind to one another with high specificity, providing a practical “programming language” for directing assembly. Yet even with major advances in molecularly programmable approaches, translating an arbitrary three-dimensional (3D) superlattice into precise, experimentally feasible assembly instructions remains a central challenge. Researchers at the Center for Functional Nanomaterials and Columbia University address this gap with MOSES, a symmetry-mapping bond-assignment algorithm for inverse design of 3D lattices self-assembled from voxels bearing directional, addressable bonds and capable of carrying nanocargo. By exploiting lattice symmetries while explicitly enforcing experimentally relevant DNA binding rules and restrictions, MOSES minimizes the number of distinct DNA-based voxels required, reducing the information that must be encoded to program assembly. The researchers demonstrated the approach on selected examples including nanoscale analogs of zinc blende (ZnS), the cubic Laves phase (MgCu2), and a lattice built from an arbitrarily prescribed motif (the letter H). Most importantly, MOSES produces clear, machine-readable “recipes” that automated lab systems can follow—helping turn design ideas into real experiments faster and making it easier to repeat, refine, and scale up complex 3D nanomaterial assembly.
Publication Reference
J.S. Kahn, D.C. Redeker, A. Michelson, A. Tkachenko, S. Hong, B. Minevich, and O. Gang. ACS Nano 2025 19 (15), 14795-14807.
DOI: 10.1021/acsnano.4c17408
OSTI: https://www.osti.gov/biblio/2555913
Acknowledgment of Support
The work was supported by the U.S. Department of Energy, Office of Basic Energy Sciences, Grant DE-SC0008772 and by U.S. Department of Defense, Army Research Office, Grant W911NF-22-2-0246. This research used resources of the Center for Functional Nanomaterials, which are U.S. DOE Office of Science Facilities at Brookhaven National Laboratory under contract no. DE-SC0012704. The data are available upon request.
2026-22836 | INT/EXT | Newsroom